What are bacteria?

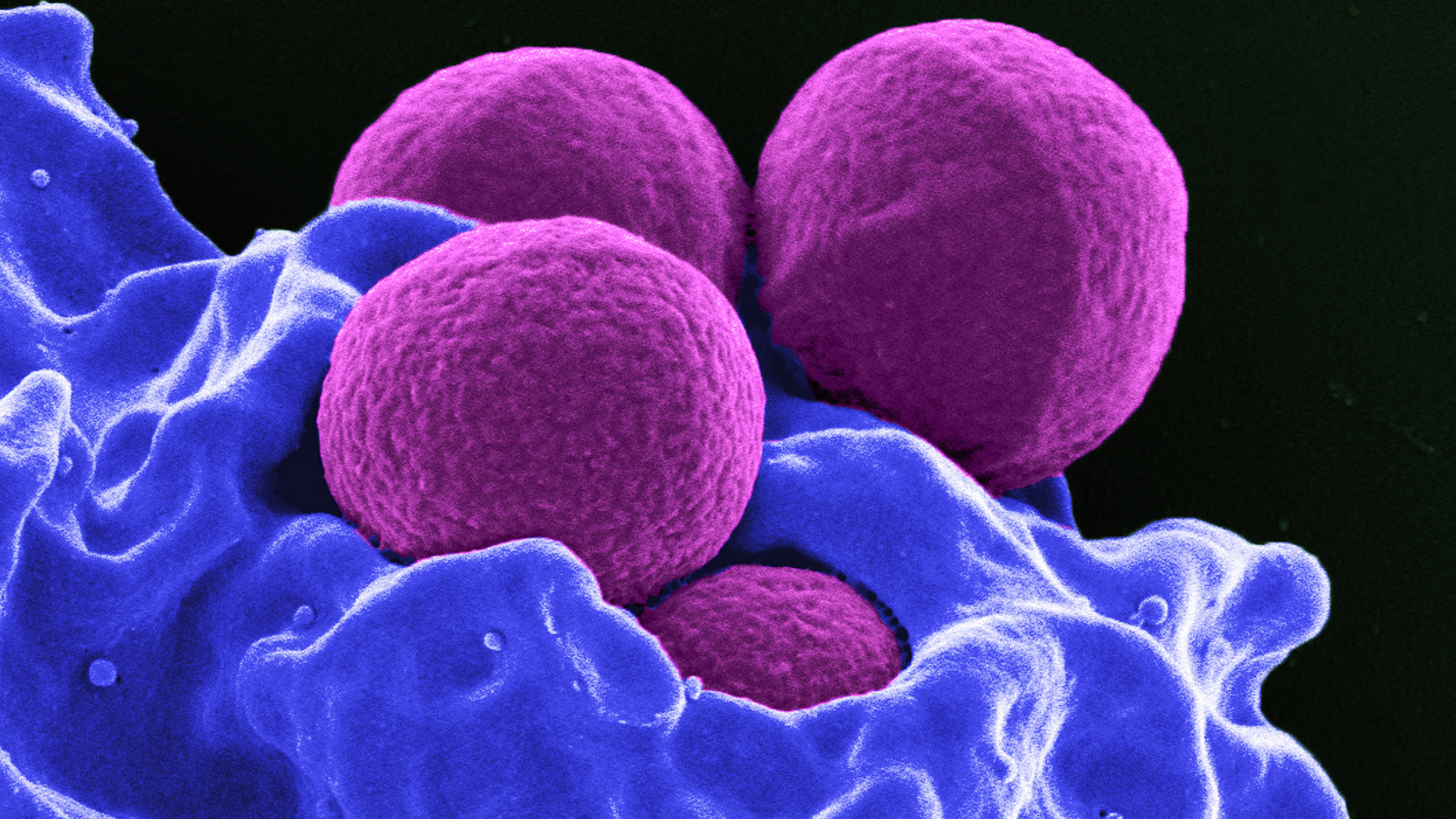
This colorized image (a scanning-electron micrograph) shows four spherical methicillin-resistant Staphylococcus aureus (MRSA) bacteria (purple) in the process of being “ingested” by a human neutrophil white blood cell (blue).
Bacteria are single-celled organisms that are pretty much everywhere: in the ground, in the ocean, on your hands and in your gut. While some are harmful, most are not — and some are even beneficial to human health. In many cases, humans live in symbiosis with bacteria, maintaining a mutually beneficial relationship without even knowing it.
So let's demystify this diverse group of single-celled organisms. Here is an overview of what bacteria are, what they do and which ones to watch out for.
What are bacteria?
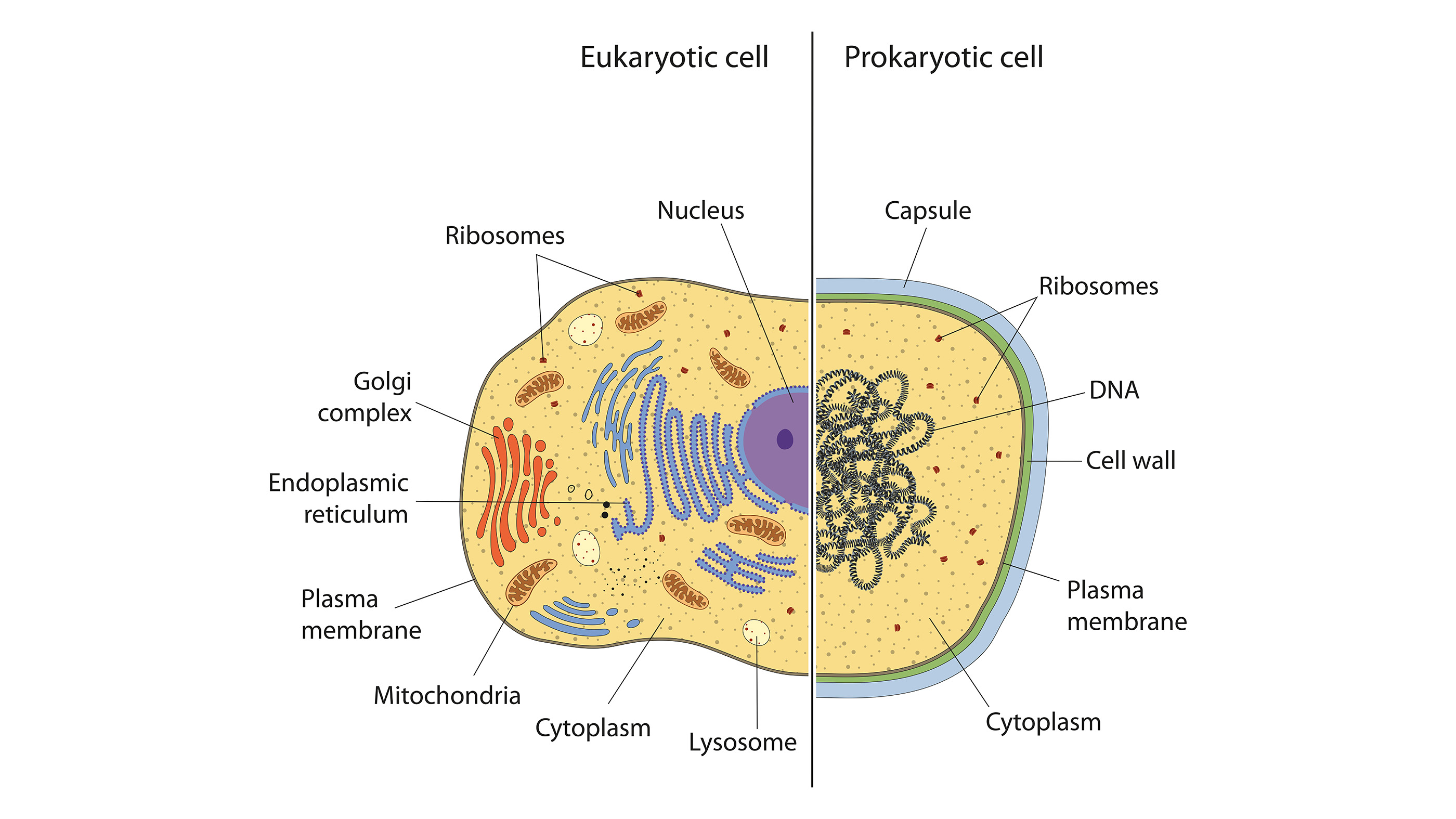
Bacteria are single-celled organisms with a unique internal structure. Humans and other multicellular organisms are eukaryotes, which means our cells have distinct nuclei bound with a membrane. Bacteria are prokaryotes, meaning they don't have organized nuclei or any other membrane-bound organelles.
Bacterial DNA floats freely within bacterial cells in a twisted, thread-like mass called the nucleoid. Some also have separate, circular pieces of DNA called plasmids. According to the Microbiology Society, plasmids often contain genes that give bacteria a survival edge, such as genes conveying antibiotic resistance.
Bacteria shouldn't be confused with the other major group of prokaryotes called archaea. Archaea are also single-celled organisms, but the two groups differ in the kinds of molecules they use to build their cell walls and in the metabolic processes they use.
Structure of bacteria
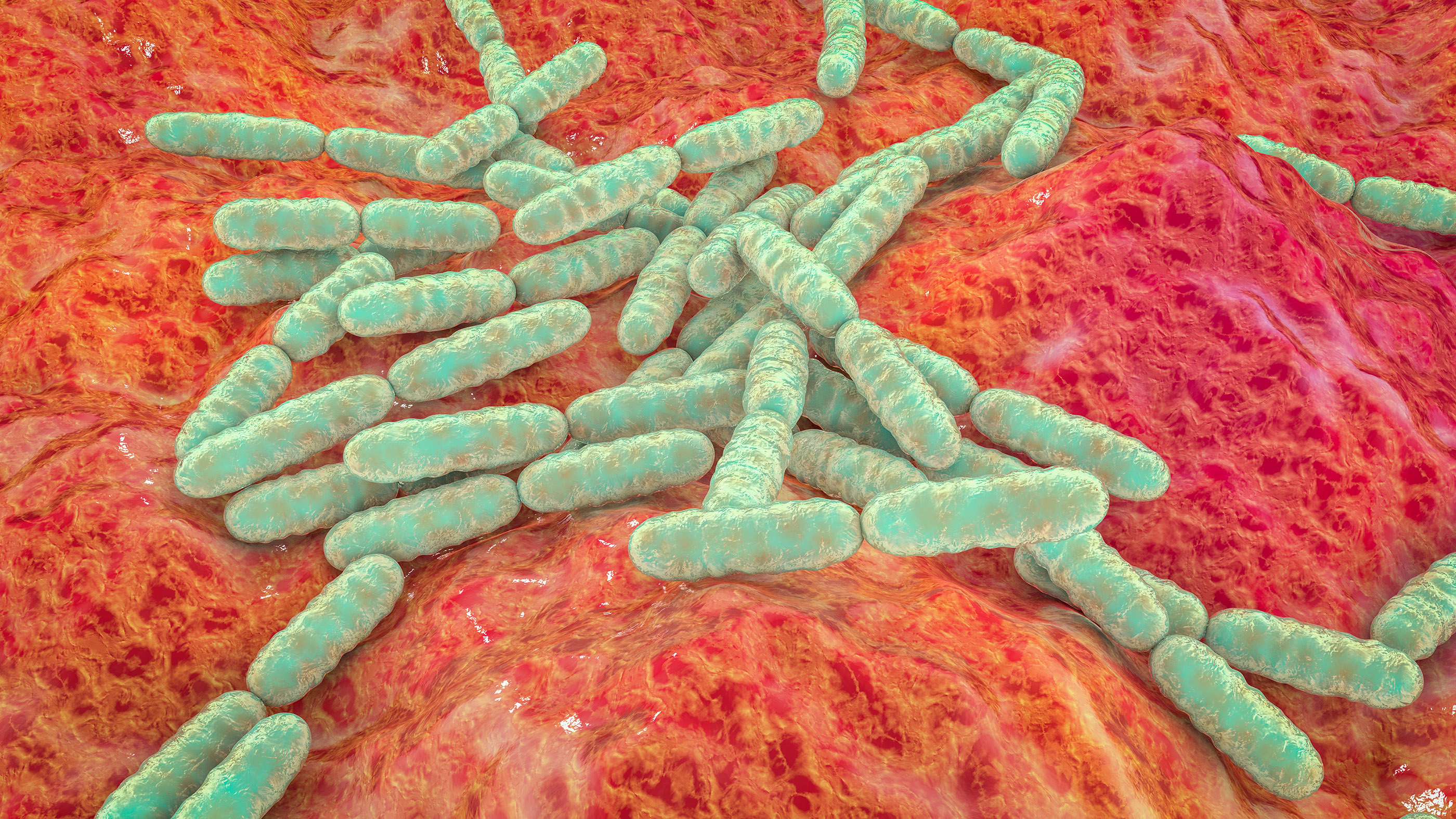
Milk-curdling Lactobacillus acidophilus are bacilli bacteria, meaning they are cylindrical-shaped.
Bacteria come in five basic shapes: spherical, cylindrical, comma-shaped, corkscrew and spiral. The scientific names for these shapes are cocci (round), bacilli (cylindrical), vibrios (comma-shaped), spirochaetes (corkscrew) and spirilla (spiral). The shapes and configurations of bacteria are often reflected in their names. For example, the milk-curdling Lactobacillus acidophilus are bacilli, and pneumonia-causing Streptococcus pneumoniae are a chain of cocci.
Bacterial cells are generally surrounded by an outer cell wall and an inner cell membrane. Certain bacteria, like the mycoplasmas, do not have a cell wall at all. Some bacteria may even have a third, outermost protective layer, called the capsule. Whip-like extensions often cover the surfaces of bacteria — long ones, called flagella, or short ones, called pili — and help bacteria move around and attach to a host.
Get the world’s most fascinating discoveries delivered straight to your inbox.
Bacteria can be classified by the composition of their cell walls using a test called the Gram stain, according to the Science Education Resource Center at Carleton College. The test stains Gram-positive bacteria, or bacteria that do not have an outer membrane. Gram-negative bacteria, which do have an outer membrane, don't pick up the stain. For example, S. pneumoniae is a Gram-positive bacterium, but Escherichia coli, which can cause food poisoning, and Vibrio cholerae, which causes cholera, are Gram-negative bacteria.
Delving beneath the cell wall and membrane, bacteria contain cytoplasm, a solution of mostly water and salts. Within the cytoplasm float the nucleoid, plasmids and tiny protein factories called ribosomes, which are the sites where the cell's genetic instructions are translated into the cell's products. Some antibiotics, like tetracycline, target bacterial ribosomes to prevent them from synthesizing proteins, thus dooming the cell.
The cytoplasm of some bacteria may also have little pockets, called inclusions, where nutrients are stored for lean times. Photosynthetic bacteria, which generate energy from sunlight, may have structures called chromatophores spread throughout their cytoplasm. These chromatophores hold pigments used in photosynthesis.
How do bacteria eat and reproduce?
As some of the oldest life-forms on Earth, bacteria have evolved a dizzying number of ways to survive. Some bacteria are photosynthetic, while others are master decomposers, breaking down rotting and decaying organic material into nutrients. Some enter symbiotic, or mutually beneficial, relationships with a host (more on this later).
Most bacteria multiply by a process called binary fission, according to the Cornell University College of Agriculture and Life Sciences. In this process, a single bacterial cell, called the "parent," makes a copy of its DNA and grows larger by doubling its cellular content. The cell then splits apart, pushing the duplicated material out and creating two identical "daughter" cells.
Some types of bacteria, such as cyanobacteria and firmicutes, reproduce via budding. In this case, the daughter cell grows as an offshoot of the parent. It starts off as a small nub, grows until it is the same size as its parent and then splits off.
After binary fission or budding, the DNA found in parents and offspring is exactly the same. Therefore, bacterial cells introduce variation into their genetic material by integrating additional DNA, often from their surroundings, into their genome. This is known as horizontal gene transfer, according to the San Diego State University College of Sciences. The resulting genetic variation ensures that bacteria can adapt and survive as their environment changes, Live Science previously reported.
There are three ways horizontal gene transfer occurs: transformation, transduction and conjugation.
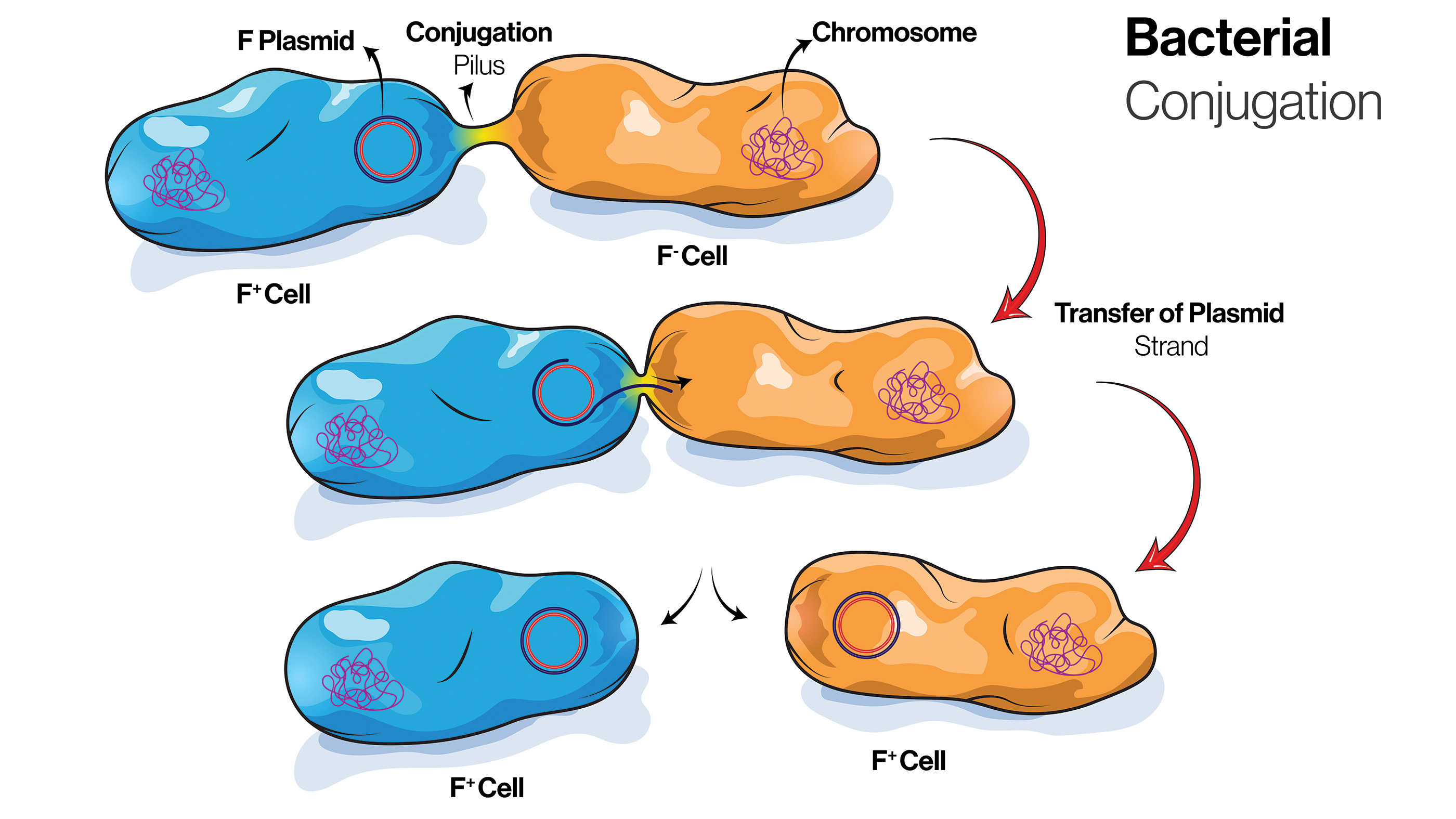
This diagram of shows the stages of bacterial conjugation.
Transformation is the most common process of horizontal gene transfer and occurs when a bacterium absorbs short DNA fragments from the environment through its cell membrane. (The DNA fragments are released into the environment by other bacteria.) To undergo transformation, a bacterium must be in a state known as competence. This usually happens when nutrients are scarce or when the density of a bacterial colony is high. In these circumstances, it might be evolutionarily advantageous to try out some new DNA.
Transduction occurs when a virus picks up DNA from one bacterium and infects another bacterium, inserting the new gene sequence. Conjunction happens when bacteria make direct contact. A donor cell sprouts a tube-like appendage, called a pilus, and directly passes DNA to a recipient cell. This happens with E. coli, in which some individual cells carry a special type of plasmid known as the fertility factor, or F factor, according to "Modern Genetic Analysis" (W. H. Freeman and Company, 1999). These F factor cells can donate DNA to cells that are F factor-negative. The third type of transfer, called conjugation, aids in the spread of antibiotic-resistance genes.
How are bacteria beneficial to health?
Many bacteria are beneficial to humans. We harness their power to curdle milk into yogurt and ferment cabbage into kimchi. Some species even do their work inside us. According to the Microbiology Society, there are about 10 times as many bacterial cells as human cells inside a person's body, and many of these live in the digestive tract. These bacteria get a constant stream of nutrients from the human gut. In return, they help break down foods that human digestive enzymes can't. Bacteroides thetaiotaomicron, for example, helps break down complex carbohydrates. L. acidophilus breaks down sugars in milk and creates byproducts such as lactic acid and hydrogen peroxide, according to the Mount Sinai health library; these byproducts make the intestines less hospitable for harmful bacteria.
Bacteria on the skin can also produce byproducts that keep harmful bacteria away, according to a 2018 paper in the journal Nature Reviews Microbiology. The benign bacterium Corynebacterium accolens, for example, inhibits the growth of the pneumonia-causing S. pneumoniae.
Some skin bacteria can be both helpful and harmful. Staphylococcus epidermidis is a spherical bacterium that commonly colonizes the skin but can cause infection if it gets inside the human body. However, S. epidermidis also produces proteins that inhibit the growth of its more virulent relative, Staphylococcus aureus. S. aureus also causes infections when it gets past the barrier of the skin, but they're typically much more serious than S. epidermidis infections.
How are bacteria harmful to health?
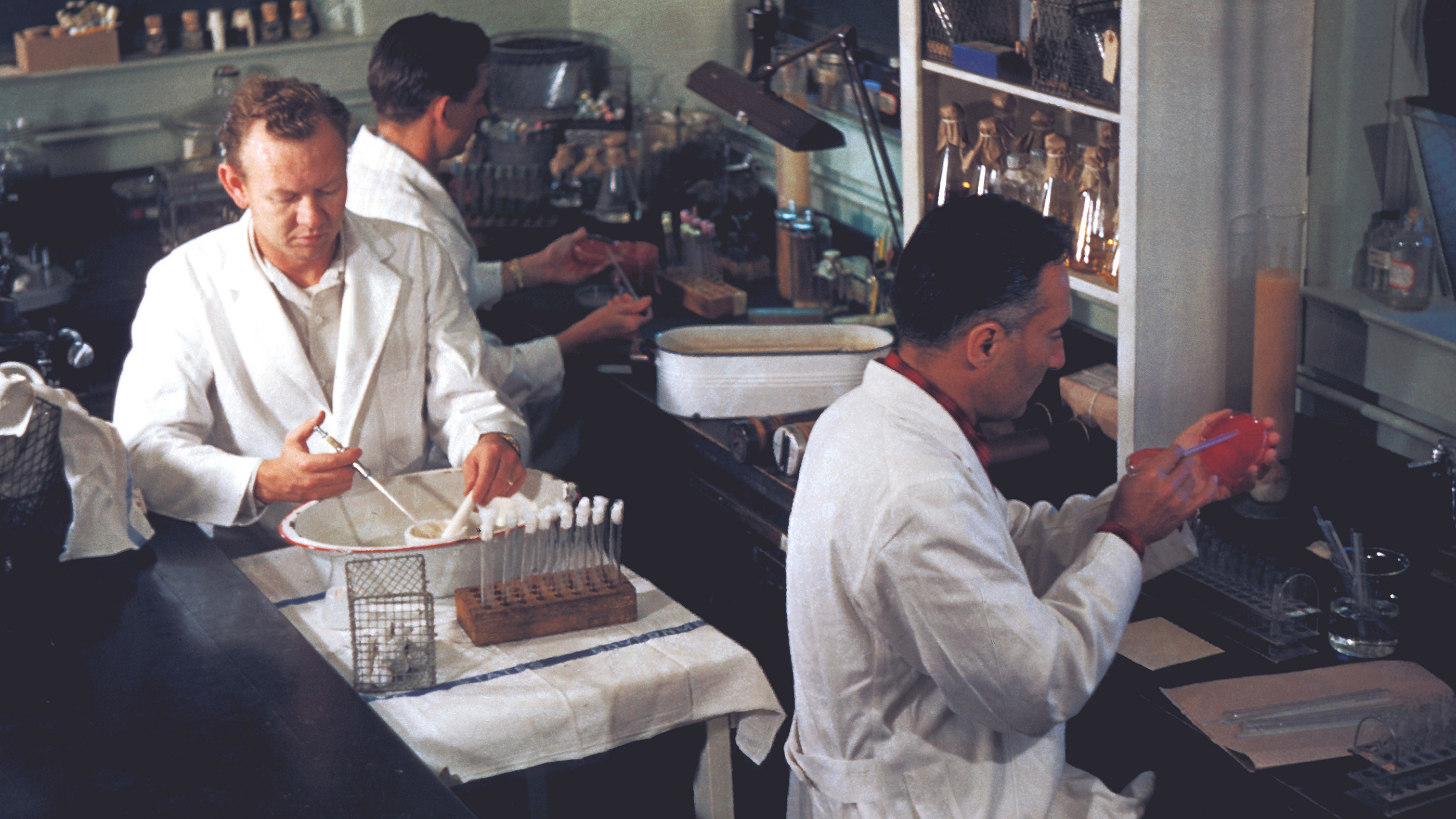

Technicians in a bacteriology laboratory in San Francisco, isolate the bacteria Yersinia pestis during a plague study in 1965.
Some bacteria, like S. aureus, live in relative peace with humans most of the time; about 30% of people carry S. aureus in their noses, according to the Centers for Disease Control and Prevention (CDC). But when these bacteria find their way into the body, especially in people who are immunocompromised, they can cause fatal infections. Staph infections can cause sepsis (full-body inflammation in response to infection), pneumonia, endocarditis (inflammation of the heart and heart valves) and osteomyelitis (inflammation of the bone), according to the CDC.
Other bacteria are almost always harmful to humans. Cholera, a diarrheal disease that kills about 95,000 people worldwide each year, is caused by V. cholerae. The bacterium Yersinia pestis, spread by fleas that bite rodents, was responsible for the Black Death. And Bacillus anthracis can form almost indestructible anthrax spores that lurk in soil and can cause deadly disease if inhaled or consumed.
Some of the most common problematic bacteria infect people through spoiled food. Salmonella bacteria cause an illness called salmonellosis, marked by diarrhea, stomach cramping and fever. Though most people recover after four to seven days, salmonellosis can be serious, and even fatal, in both young children and older people, according to the CDC.
E. coli, another bacterium responsible for food poisoning, often spreads through contaminated food and water. While many strains live harmlessly in human intestines, others cause diarrheal illness. Like salmonellosis, E. coli diarrhea is typically deeply unpleasant but brief, though 5% to 10% of people develop a kidney complication called hemolytic uremic syndrome, which can be life-threatening, according to the CDC.
Another common bacterium that can be harmful to people is Helicobacter pylori. About half of people carry these bacteria in their stomachs, according to the Mayo Clinic. Most people never show any ill effects of this infection, but in some cases, the bacteria cause peptic ulcers, or painful sores in the lining of the stomach. It's not entirely clear how the bacteria spread, but risk factors include crowded livi
What is bacterial vaginosis?
Bacterial vaginosis is a condition in which anaerobic bacteria (bacteria that do not use oxygen in their metabolism) overtake Lactobacillus, a type of beneficial bacteria, in the vagina. Symptoms include vaginal itching, gray or green discharge, a fishy odor and pain during urination, according to the Mayo Clinic.
Bacterial vaginosis is common: A national study conducted between 2001 and 2004 found that 29% of women randomly tested for the bacteria that cause the condition had it, corresponding to about 21 million women in the U.S. affected at any given time. (Only about 15% of the women who tested positive had symptoms.)
It's not clear what causes bacterial vaginosis, according to the Mayo Clinic. Some people are likely susceptible because their vaginal environment is not as comfortable for the Lactobacillus bacteria that make up a healthy vaginal microflora. Douching or having sex with a new partner or with multiple sexual partners can be a risk factor, possibly because these activities disrupt the usual bacterial communities in the vagina. Bacteria commonly associated with bacterial vaginosis include Gardnerella vaginalis, Prevotella species, Mobiluncus species and Atopobium vaginae, according to the CDC.
If untreated, bacterial vaginosis is a risk factor for preterm birth and can make a person more susceptible to contracting sexually transmitted infections. Bacterial vaginosis can be treated with antibiotics.
Antibiotic resistance
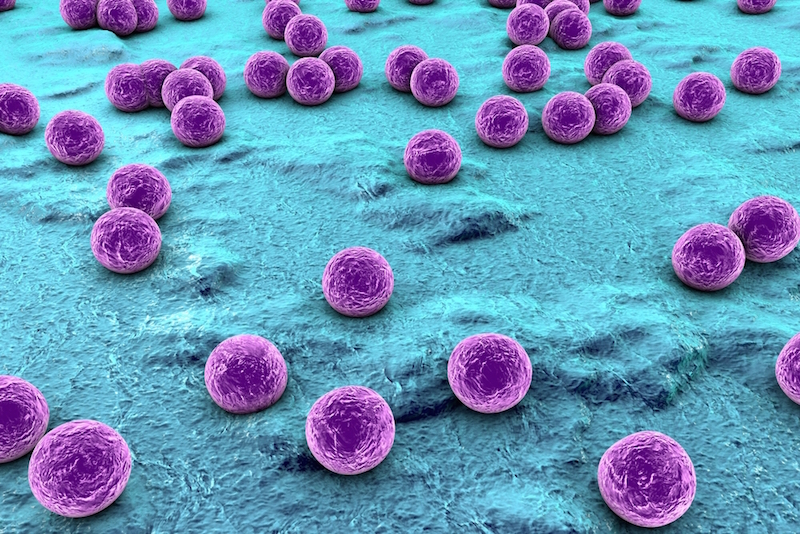
Antibiotics are typically used to treat bacterial infections. However, in recent years, improper and unnecessary use of antibiotics has promoted the spread of several strains of antibiotic-resistant bacteria.
In cases of antibiotic resistance, the infectious bacteria are no longer susceptible to previously effective antibiotics. According to the CDC, at least 2 million people in the U.S. are infected with antibiotic-resistant bacteria every year, leading to t23,000 deaths annually.
"Pretty much any infection you can think of now has been identified as being associated with some level of resistance," said Dr. Christopher Crnich, an infectious-disease physician and hospital epidemiologist at the University of Wisconsin Hospitals and William S. Middleton Memorial Veterans Hospital. "There's very few infections that we now treat where infections caused by resistant bacteria is not a clinical concern."
MRSA, for example, is one of the more notorious antibiotic-resistant bacterial strains; it resists methicillin and other antibiotics used to treat Staphylococcus infections, which are acquired primarily through skin contact. MRSA infections occur in health care settings such as hospitals and nursing homes, where it can lead to pneumonia or bloodstream infections. MRSA also spreads in communities, especially in situations where there is a lot of exposed skin, other physical contact and the use of shared equipment — for example, among athletes, in tattoo parlors, and in day care facilities and schools, according to the CDC. Community-acquired MRSA most often causes serious skin infections.
An important facet of combating antibiotic resistance is to be careful about their use. "It's so important for us to use antibiotics intelligently," Crnich told Live Science. "You only want to use an antibiotic when you have a clear-cut bacterial infection."
Additional resources:
- Read more about the life history and ecology of bacteria from the University of California, Berkeley.
- Watch: Bacteria: Energy Producers of the Future? From the National Science Foundation.
- Learn how bakers and their bread are a microbial match, from NPR.
This article was updated on Oct. 14, 2021, by Live Science Contributor Stephanie Pappas.
Aparna Vidyasagar is a freelance science journalist who specializes in health and life sciences. Aparna has written for a number of publications, including New Scientist, Science, PBS SoCal, Mental Floss, and several others. Aparna has a doctorate in Cellular and Molecular Pathology from the University of Wisconsin-Madison, and also received a master’s degree and bachelor’s degree from the same university.

 Live Science Plus
Live Science Plus




