
Ctenophores Semaphore Information About Earliest Animals

This article was originally published at The Conversation. The publication contributed the article to LiveScience's Expert Voices: Op-Ed & Insights.
Most of us have never heard of Ctenophores, or comb jellies, but this is about to change. In a publication out today in Science, a team of researchers in the computational genomics unit at the National Institutes of Health in Maryland report that Ctenophora are the most ancient multicellular animals. This was a spot previously held by sponges.
To understand the implications of this finding, we have to remember that multicellularity was a big step in evolution that occurred over 550 million years ago. At the time there was an explosion of forms, as life explored the limitations and possibilities of having a body made up of different types of cells.
The arrangement of cell types continues to be the basis of animal classification. Sponges were the obvious choice for the first experiment in cellularity, as they have no nervous system, few cell types and no organisation of tissues.
Studying evolutionary patterns relies on comparing genetic information. Animals that appear superficially similar (such as jellyfish and comb jellies) can be quite different at a genetic level. Modern taxonomy has embraced barcoding, which uses the DNA sequence of a single gene to distinguish between closely related species. But one gene never tells the whole story, and when looking back to the beginning of metazoan evolution, even multiple genes can lead us astray.
The breakthrough in today’s paper is the sequencing of the entire genome of a Ctenophore known as the sea walnut (Mnemiopsis leidyi). This was then compared with the entire genome of organisms from the other main groups at the base of the animal tree: Porifera (sponges), Cnidaria (jellyfish and anenomes) and Placozoa (there is no common name for Placozoa).
Together these animals comprise the non-bilateria, which only makes sense when you realise that most of the animals we know are Bilaterians: insects and fish and people and dogs all have bilateral symmetry.
Get the world’s most fascinating discoveries delivered straight to your inbox.
It now appears that the closest relative of Bilaterians are jellyfish, while the most ancient animals are the comb jellies.
What is a comb jelly?
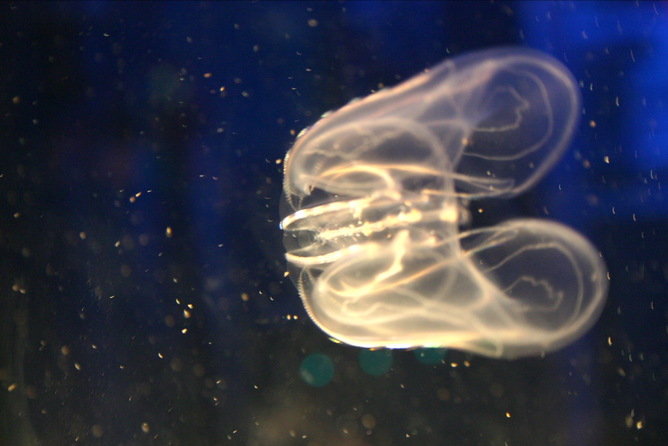
Ctenophores are delicate, translucent creatures. They have eight rows of comb plates with cilia that provide them with locomotion. They are carnivorous, hermaphroditic marine creatures that do not sting.
The sea walnut (M. leidyi) is native to the western Atlantic but has been introduced to the Black, Caspian and North Seas where it has caused serious environmental and economic damage by eating native zooplankton and fish. Apparently, it glows blue green when disturbed.
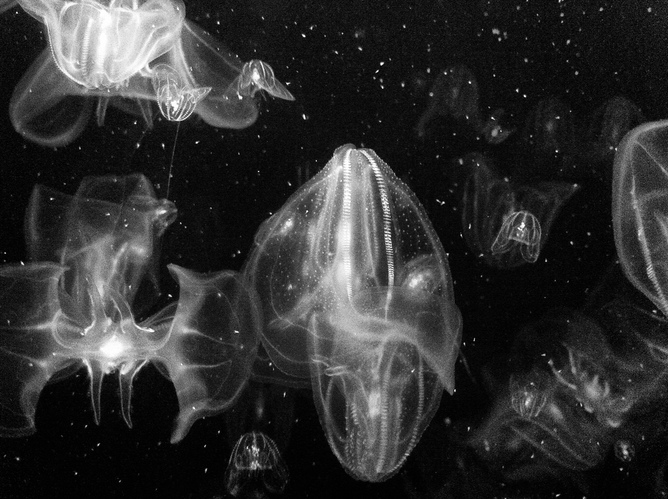
In terms of cellular arrangements, Ctenophores have a nervous system and all three major cell types (endoderm, ectoderm and mesoderm).
Sponges, by contrast, have no cell types and no nervous system. No wonder we thought sponges were the more primitive organism.
What did we learn about metazoan genomes?
The sea walnut genome contained 16,548 protein coding genes, 44% of which shared homology - a type of ancestry - with non-Ctenophores. Comparing these genomes with those of the other major animal groups allowed the authors to reject several hypotheses about early animal evolution.
The amount of computational effort to achieve these aims is hard to overstate, with an average run of 205 days for the Bayesian computer analyses alone.
We can study cell types and body plans in two ways: by examining the tissues themselves or by comparing genetic pathways available to create certain tissues. The latter approach has been used quite effectively in this study.
For example, Ctenophores have a nervous system and sponges do not, but sponges do have the genes required for nervous system development and function. This means that the ancestor of all animals may have had quite an advanced nervous system and these structures (but not their genes) were lost in the lineage that led to sponges.
Another major finding concerned the development of the main cell types in early animals. Embryonic cell layers develop into specific types of tissues. Ectoderm formed the skin and nervous system, endoderm formed the gut and mesoderm provided the muscle. Until recently, it was thought that non-bilaterians lacked the mesoderm layer.
Ctenophores, however, do have a third cell layer called a mesoglea which acts like muscle. The genes that support mesoglea development are completely unique and sufficiently different from bilaterian mesoderm to suggest an independently evolved three-layer system in these seemingly simple forms.

Ancient does not equal primitive
As sea walnuts glow when disturbed, so does this study shed light on some interesting assumptions about animal evolution.
The first experiments in multicellularity were not simple collections of cells without structures or communication. Genes responsible for cell signalling were present even before the evolution of multicellular animals. This suggests single celled organisms were communicating with each other before they decided to organise themselves into bodies with different types of cells.
Second, the three cell layers of well-known animals including ourselves is not unique nor is it a latecomer to animal evolution. The earliest multicellular animals evolved their own form of mesoderm independently, with unique genes allowing sophisticated biological organisation.
Third, the ancestor of all animals had a nervous system which coordinated bodily functions. The nervous system was subsequently lost in the lineages that led to Porifera and Placozoa, but survived in the Cnidarians and the Bilaterians.
Finally and most profoundly, the shape of the evolutionary tree of all animals has taken on a new shape. The earliest branch of the animal tree belongs to Ctenophora, now confirmed to be the sister lineage to all other animals.
So don’t confuse comb jellies with jellyfish.
I think of the Ctenophores as a semaphore, signalling some profound truths to us (in a blue green glow) across the vastness of time about animal origins and biological organisation.
Susan Lawler has received funding from the ARC in the past.
This article was originally published at The Conversation. Read the original article. The views expressed are those of the author and do not necessarily reflect the views of the publisher. This version of the article was originally published on LiveScience.

 Live Science Plus
Live Science Plus





