How does CRISPR work?
CRISPR is a versatile tool for editing genomes and has recently been approved as a gene therapy treatment for certain blood disorders.


CRISPR, short for CRISPR-Cas9, is a genome-editing tool that allows scientists to precisely cut and modify DNA sequences. It has revolutionized the study of genes, helped to enhance crops and improved health care.
The gene-editing system was originally discovered in bacteria, where it limits infections by clipping viral DNA. Then, in Nobel prize-winning work, this bacterial defense apparatus was co-opted by scientists to devise a new approach to genome editing.
"It's really the simplicity, the cost and the ease of use" that democratized this editing tool, Alison Van Eenennaam, a livestock geneticist at the University of California, Davis who uses CRISPR to alter the genetics of farm animals, told Live Science.
Recently, CRISPR has been approved to treat two blood disorders, and early-stage trials reveal its potential to treat inherited blindness. Here's everything you need to know about the groundbreaking technology.
What is CRISPR?
The CRISPR system includes the following major components:
CRISPR: "CRISPR" stands for "clusters of regularly interspaced short palindromic repeats." This unwieldy name describes a pattern of DNA sequences found in bacterial genomes that helps the bacteria fend off viruses.
Get the world’s most fascinating discoveries delivered straight to your inbox.
"Short palindromic repeats" refers to sequences that read the same forward and backward, like the words "kayak" and "racecar." DNA consists of two paired strands twisted around each other in a helix. A DNA palindrome thus refers to a string of DNA letters, or bases — A (adenine), C (cytosine), G guanine) and T (thymine) — that are the same when read forward on one strand and read backward on the other.
For example, if one strand reads "GATC" in one direction, these DNA bases will pair up with "CTAG" on the opposite strand because G always pairs with C and A always pairs with T. When read backward, CTAG becomes the original sequence, GATC.
These repeats are "regularly interspaced," meaning that these CRISPR regions in the genome contain an alternating pattern of palindromes with "spacer" sequences wedged between them. Bacteria co-opt the spacer sequences from the DNA of invading viruses and store them in their CRISPR regions to fight future infections.
This system is often likened to the human adaptive immune system, which similarly stores a "memory" of previous infections in order to stave off repeat encounters. Rather than using immune cells, like humans do, bacteria use CRISPR.
CRISPR RNA (crRNA) and Cas9: CRISPR DNA serves as a permanent record of past infections, but for bacteria to use these sequences to thwart viruses, they must convert them into DNA's cousin, RNA. Through a process called transcription, bacteria first copy one of the two CRISPR DNA strands into a single complementary strand of RNA; the strand is complementary in that it matches the original DNA code, except it replaces T with U (uracil). Then, the microbes chop the lengthy strand into shorter crRNA snippets, each carrying one repeat and one spacer.
The bacteria also make a second RNA molecule, called "trans-activating crRNA," or tracrRNA. This RNA contains a reversed version of the palindromic repeat on the crRNA molecule, allowing the two RNAs to bind together.
The resulting complex can then latch onto viral DNA carrying the spacer sequence, calling forth an enzyme that cuts and disables that DNA. The enzyme, called "CRISPR-associated protein 9," or Cas9, is essentially a pair of molecular scissors.
There are also other types of Cas enzymes that can be utilized in gene editing. For example, one called Cas12a produces staggered cuts in DNA, in which one strand is longer than the other at each end. DNA sequences can then be paired with the overhanging strand. Cas14 makes cuts in RNA, instead of DNA, and could be useful for temporarily altering what proteins a cell makes without making permanent edits to its genome.
Related: Meet 'Fanzor,' the 1st CRISPR-like system found in complex life
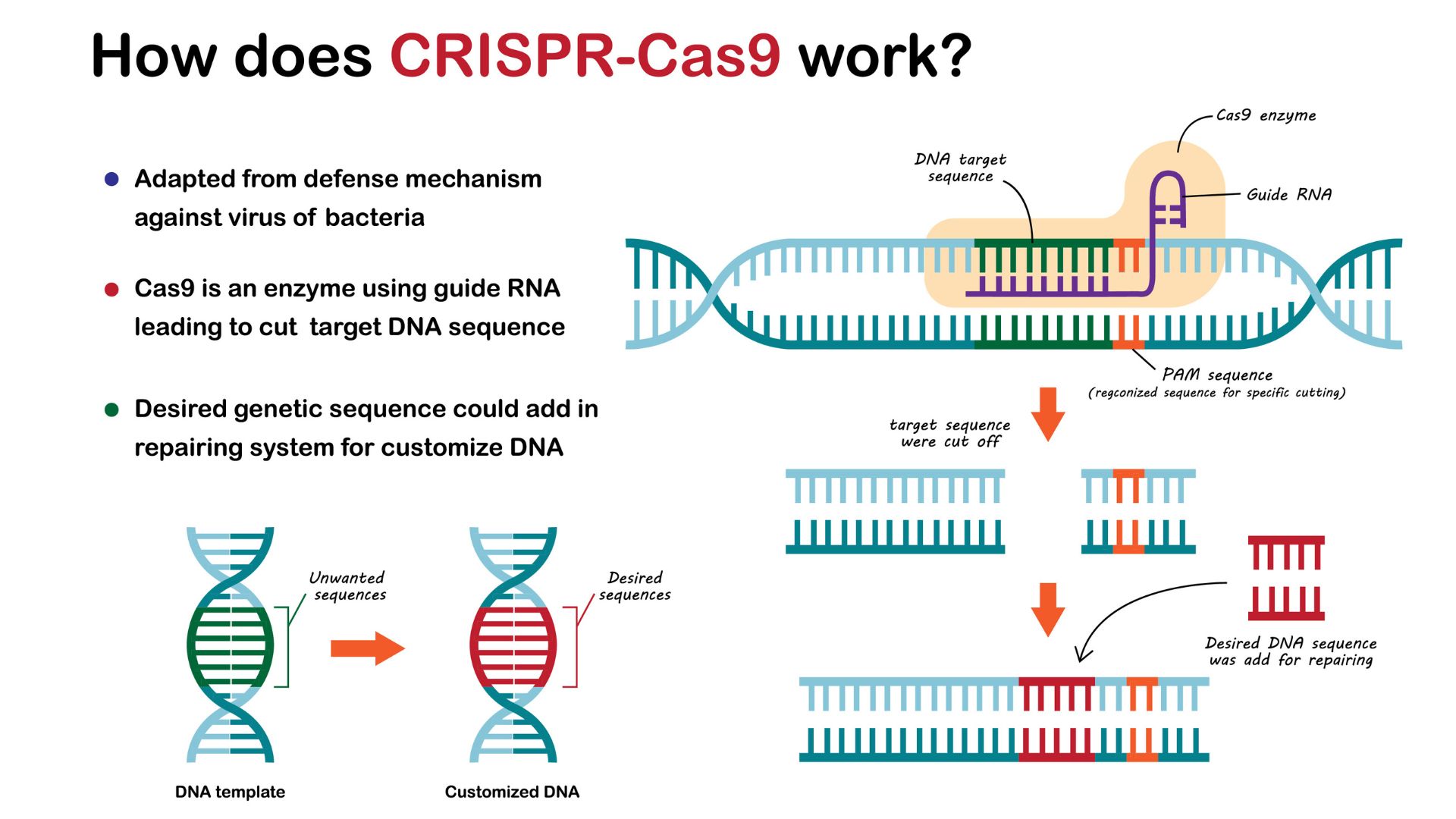
CRISPR gene-editing systems work by guiding Cas enzymes to a specific place in the genome that they then cut through.
How does CRISPR edit DNA?
Researchers have taken advantage of the CRISPR system's ability to make precise cuts in DNA. By adapting CRISPR to make desirable genome edits in any cell type, researchers can alter genes or DNA sequences that regulate genes' activity, changing their function or expression.
To simplify the system, scientists combined the crRNA and tracrRNA molecules described in the previous section into a single molecule called "guide RNA."
"All you need to do to target a new sequence is alter the guide," Van Eenennaam said, which is cheap and quick to do. In contrast, other genome-editing techniques require the time-consuming and expensive design of a lab-made protein that targets a sequence of interest.
The guide RNA is paired with a Cas9 enzyme to make edits in the genome. Once the RNA binds to the desired sequence, the enzyme swoops in and cuts both strands of DNA. In response, the cell attempts to glue the strands back together, but it uses a fault-ridden process that often introduces mutations. For instance, it may add a few extra letters. This change often deactivates the gene, making CRISPR editing a simple strategy to switch off genes.
Scientists have also modified the Cas9 enzyme to make other types of edits. By disabling Cas9's genetic scissors and then fusing this "dead Cas9" to another enzyme, they can rig the machinery to alter single bases, converting a C into a T, for example. This CRISPR formulation is called "base editing," and it allows researchers to make fine changes that alter the structure of the product encoded by the gene, whether that's a protein or RNA.
Dead Cas9 has also been paired with enzymes that activate or silence genes to tune their activity. Dead Cas9 has also been fused to fluorescent proteins, which light up when the guide RNA binds to a specific stretch of DNA, essentially revealing its post code in the cell.
Related: Could CRISPR cure HIV someday?
Who discovered CRISPR?
CRISPR's history goes back to 1987, when Yoshizumi Ishino and colleagues at Osaka University in Japan first reported the unusually repetitive sequences in Escherichia coli, a well-known bacterium. At the time, the scientists didn't know how these clusters were related to bacterial defense.
In the 1990s, these clusters drew the attention of more scientists when Francisco Mojica (who coined the term "CRISPR") and his team at the University of Alicante in Spain spotted them in 20 other bacterial genomes, suggesting they had widespread importance in bacteria.
In 2005, Alexander Bolotin and colleagues at the French National Institute for Agricultural Research came across genes for Cas enzymes located near the CRISPR region of a genome. Shortly after, Eugene Koonin's group at the National Institutes of Health revealed that the spacer sequences matched viral DNA, leading researchers to connect CRISPR with bacterial immunity.
Emmanuelle Charpentier, of the Max Planck Institute in Germany, and Jennifer Doudna, of the University of California, Berkeley, later adapted CRISPR for genome editing. Their work led them to share the 2020 Nobel Prize in chemistry.
Shortly after the groundbreaking publication of Charpentier's and Doudna's work, Virginijus Šikšnys of the Vilnius University Institute of Biotechnology and his colleagues also demonstrated how CRISPR could be used in gene editing. Feng Zhang's group at the Broad Institute later developed other CRISPR systems into gene-editing tools, including an RNA-editing system involving an enzyme called Cas13.
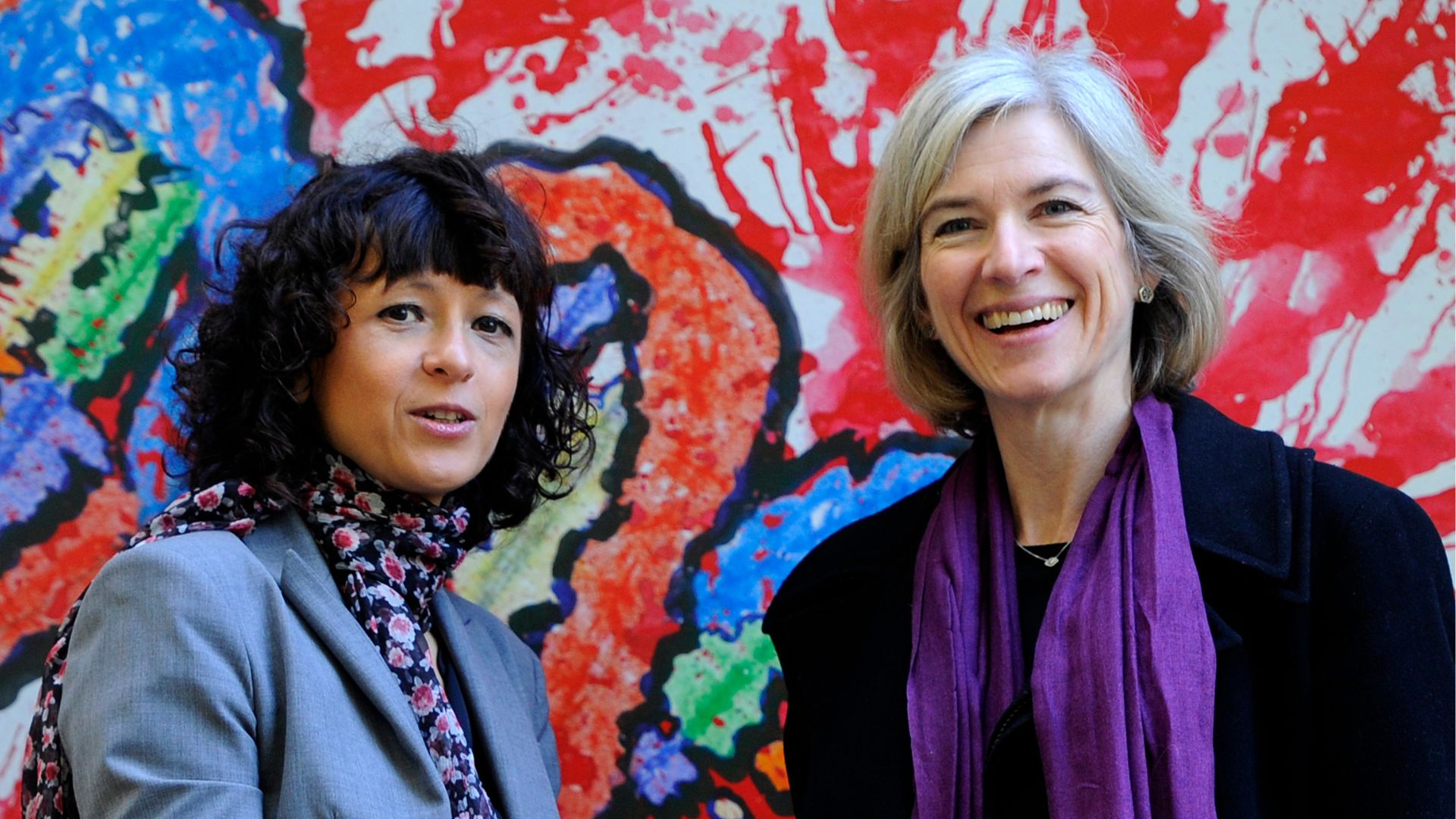

Emmanuelle Charpentier (left) and Jennifer Doudna received a Nobel Prize for their pioneering work with CRISPR.
How has CRISPR been used in people? What about in plants and animals?
CRISPR has been used to correct genetic disorders, such as cystic fibrosis and cataracts, in laboratory-grown cells and in lab animals. It has also shown recent success as a treatment for other conditions in human trials. Notably, the U.K. and the U.S. have both approved a CRISPR-based gene therapy called Casgevy for two blood disorders: sickle cell disease and beta thalassemia. This is the first CRISPR-based therapy to ever be approved.
Casgevy works by cutting and disabling the gene BCL11A, which controls the switch from fetal hemoglobin to adult hemoglobin shortly after birth.
The fetal version binds more strongly to oxygen, allowing a fetus to gather enough oxygen from its mother's bloodstream. The adult version normally takes over after birth, once oxygen can be obtained through breathing. However, in sickle cell disease and beta thalassemia, people have faulty versions of the adult gene. Casgevy reverses the switch to adult hemoglobin so that patients can continue using their fetal hemoglobin gene instead.
One form of inherited blindness may be among the next disorders treated using CRISPR. An early-stage trial tested injecting CRISPR components into the eye and suggested the approach to be safe and effective mode. The base-editing form of CRISPR also showed promising results in lowering cholesterol levels during a small trial.
Beyond health care, CRISPR editing has been used to enhance at least 41 food crops, including rice and wheat, by improving their palatability, nutritional value and resistance to disease. It's also been used to edit the genes of pigs whose organs are then harvested for human transplant operations.
In addition, Van Eenennaam uses CRISPR in proof-of-concept experiments to endow farmed animals with desirable traits. For example, she enhances meat yields in cattle so farmers can rear fewer livestock and thus limit their environmental impact.
What are potential dangers and downsides of CRISPR?
This is what the CRISPR system looks like as it slices through DNA.
CRISPR is a versatile and powerful genome editing tool, but at this juncture, it has some limitations and risks and also raises ethical quandaries.
For instance, in regards to treating genetic disorders, some people embrace their conditions and don't regard them as disorders, Van Eenennaam said. For instance, "should you 'cure' hereditary deafness in the offspring of a deaf couple who don't believe deafness is a bad thing?" she questioned.
Regarding CRISPR's limitations, it can introduce "off-target effects" if the Cas9 enzyme cuts DNA at unintended places in the genome. This might occur if researchers don't customize the guide RNA sequence for a unique DNA target but instead target a common sequence found in a family of genes. Such off-target effects could have negative outcomes on health. For example, if the guide RNA matches a gene that suppresses tumor growth, there is a risk that disabling it could turn the cell cancerous.
Another issue is that CRISPR editing is not 100% efficient, so only a proportion of the targeted cells undergo the desired genetic change. This means that, in some scenarios, unedited cells might avoid harmful off-target effects and thus fare better than edited cells and eventually outnumber them. Researchers recently found that edited blood stem cells can die out overtime, suggesting blood-disorder treatments may become less effective in the long haul.
These risks also pose ethical considerations regarding the use of CRISPR in livestock. Van Eenennaam typically uses "germ-line editing" in livestock, which involves targeting sex cells, like eggs and sperm, or fertilized eggs. This makes CRISPR edits heritable between an animal and its offspring. Van Eenennaam's group doesn't insert newly engineered genes into cattle, but rather, they transfer existing, desirable genes from one cow into another.
Related: Gene therapy: What is it and how does it work?
RELATED STORIES
There is a chance that off-target edits could impair the animals' health. But Van Eenennaam argues these concerns are often exaggerated. "There are going to be literally millions of genetic variations between two bulls" arising from natural mutations, so off-target effects are just a drop in the ocean, she explained.
Still, the U.S. Food and Drug Administration (FDA) says genome-editing of farmed animals requires ample oversight, in part because other DNA sequences often accompany the new gene inserted into the genome. These carryovers should be scrutinized to ensure they aren't hazardous to the animal or to human consumers, the FDA says. If the agency deems an edited animal to be low-risk, they can grant "enforcement discretion" that allows it and its descendants to be commercialized, Van Eenennaam noted.
This type of germ-line editing has rarely been applied to humans, except in the controversial case in which a Chinese scientist infamously generated "CRISPR babies" in violation of regulations. One big reason human germ-line editing has been avoided is that future generations cannot consent to receiving a CRISPR treatment.
Future generations also cannot consent to the possibility of harmful off-target effects, such as mutations that could predispose one to cancer. Nita Farahany, a bioethicist at Duke University, told the New York Times that editing embryos would be unethical "until we can figure out what the off-target effects are, and how we can control for them."
The National Academies of Sciences, Engineering and Medicine laid out criteria that should be met for germline editing clinical trials to go forward. The group advises restricting human germ-line editing to only genes whose mutations can lead to serious disease for which there are no other therapies.
Currently, gene therapies mainly use a technique called "somatic editing." This applies to Casgevy, for example. Somatic editing works by targeting a subset of non-sex cells in the body and thus doesn't pass any alterations down to additional generations.
"The benefit clearly outweighs any hypothetical risk" in these contexts, Van Eenennaam said, so somatic CRISPR editing is a "no-brainer" when it comes to gene therapy.
Editor's note: A new version of this article was published on July 1, 2024. The previous version had last been updated in March 2023.
Ever wonder why some people build muscle more easily than others or why freckles come out in the sun? Send us your questions about how the human body works to community@livescience.com with the subject line "Health Desk Q," and you may see your question answered on the website!

Kamal Nahas is a freelance contributor based in Oxford, U.K. His work has appeared in New Scientist, Science and The Scientist, among other outlets, and he mainly covers research on evolution, health and technology. He holds a PhD in pathology from the University of Cambridge and a master's degree in immunology from the University of Oxford. He currently works as a microscopist at the Diamond Light Source, the U.K.'s synchrotron. When he's not writing, you can find him hunting for fossils on the Jurassic Coast.

 Live Science Plus
Live Science Plus




