Scientists discover viruses that secretly rule the world's oceans
RNA viruses infect influential ocean organisms.

Thousands of mysterious viruses that were recently discovered lurking in the world's oceans may exert huge influence over the ecosystems, in part by "reprogramming" the hosts they infect, scientists reported.
The new research, published Thursday (June 9) in the journal Science, focuses on viruses that contain RNA, a molecular cousin of DNA. Examples of RNA viruses abound in human disease; for instance, coronaviruses and influenza viruses are both RNA-based. However, when it comes to the RNA viruses in the ocean, scientists are only just learning about the variety that can be found and the range of hosts they can infect.
Based on the new study, "we are certainly sure that most RNA viruses in the ocean are infecting microbial eukaryotes, so fungi and protists, and to a lesser extent, invertebrates," co-first author Guillermo Dominguez-Huerta, who was a postdoctoral scholar in viral ecology at Ohio State University (OSU) at the time of the study, told Live Science. Eukaryotes are organisms with complex cells that hold their genetic material inside a nucleus.
These viral hosts — namely fungi and protists, which include algae and amoebas — pull carbon dioxide out of the atmosphere and therefore influence how much carbon ends up stored in the ocean. By infecting these organisms, RNA viruses likely affect how carbon flows through the ocean at large, said Steven Wilhelm, the principal investigator of the Aquatic Microbial Ecology Research Group at the University of Tennessee Knoxville, who was not involved in the new study.
Related: 70,000 never-before-seen viruses found in the human gut
"Given the abundance of RNA virus particles, knowing they can do this continues to build the story of how important viruses are in the world with respect to how energy and carbon flow," Wilhelm told Live Science in an email.
(Wilhelm has collaborated with several of the study authors, including Matthew Sullivan and Alexander Culley, on projects unrelated to the new study.)
Get the world’s most fascinating discoveries delivered straight to your inbox.
Virus, virus everywhere
Earlier this year, Dominguez-Huerta and his colleagues reported finding more than 5,500 previously unidentified RNA viruses in the world's oceans.
For that study, which was published April 7 in the journal Science, the team analyzed 35,000 water samples that had been collected from 121 locations in the five oceans by the Tara Oceans Consortium, an ongoing global study examining the impact of climate change on oceans. These water samples teemed with plankton — tiny organisms that drift in the current and often serve as hosts for RNA viruses. To spot the viruses within these plankton, the researchers sifted through all the RNA in the planktons' cells to find a specific snippet of genetic code, called the RdRp gene.
"That's the only … coding sequence that is common across all RNA viruses," said Dominguez-Huerta, who currently works as a scientific consultant with a firm called Virosphaera; however, the RdRp gene is absent from cells and other kinds of viruses.
Ultimately, the team found so many RNA viruses tucked away in the plankton that they proposed doubling the number of RNA virus phyla — the broad taxonomic category just below "kingdom" — from five to 10 in order to classify them all.
From there, the researchers wanted to better understand how these viruses are distributed across the globe and what hosts they target.
The scientists determined that the viral communities could be sorted into four major zones: the Arctic, Antarctic, Temperate and Tropical Epipelagic, meaning close to the ocean surface, and Temperate and Tropical Mesopelagic, meaning about 656 to 3,280 feet (200 to 1,000 meters) underwater. Interestingly, the variety of viruses seemed highest in the polar zones, despite there being a wider variety of hosts to infect in warmer waters.
Related: Under the sea: 50 breathtaking images from our oceans
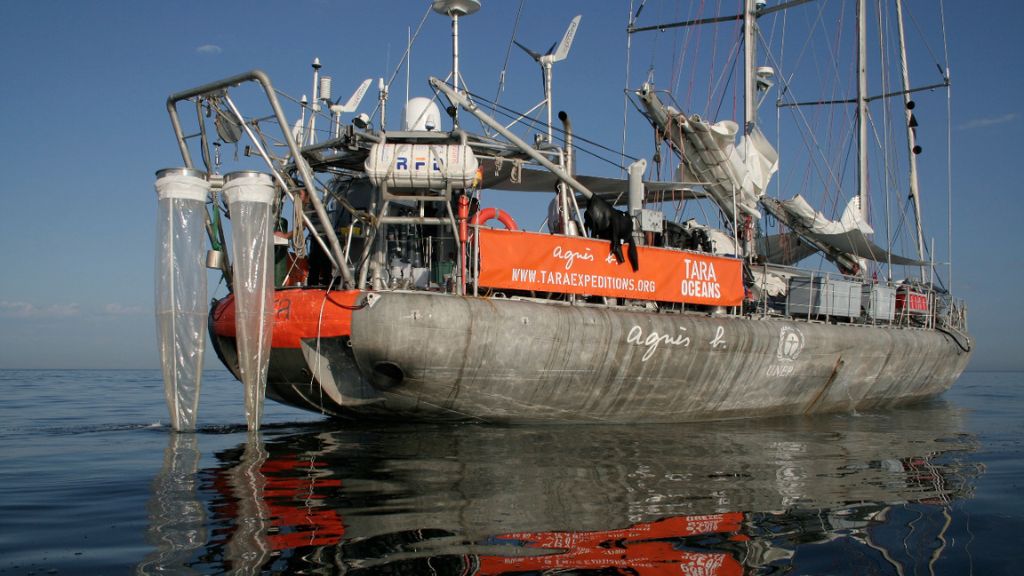

On a years-long expedition, scientists collected water samples from all the world's oceans.
"Viruses, when it comes to diversity, didn't really care about how cold the water is," said co-first author Ahmed Zayed, a research scientist in the Department of Microbiology at OSU. This finding hints that, near the poles, many viruses likely compete for the same hosts, Zayed told Live Science.
To identify these viral hosts, the team used several strategies; for instance, one method involved comparing the genomes of RNA viruses with known hosts to those of the newfound viruses, and another involved hunting for rare snippets of viral RNA in host cells' genomes, where bits of RNA can sometimes get left behind. This analysis revealed that many of the RNA viruses in the ocean infect fungi and protists, some infect invertebrates and a miniscule fraction infect bacteria.
The team also unexpectedly discovered that 95 of the viruses carried genes they'd "stolen" from their host cells, Dominguez-Huerta said. In the host, these genes help to direct metabolic processes within the cell. This discovery suggests that the viruses messed with their hosts' metabolisms in some way, likely in order to maximize the production of new virus particles, the authors concluded.
Some smaller-scale studies had hinted at this gene-swiping ability in the past, Dominguez-Huerta noted.
RELATED STORIES
After identifying what hosts the ocean viruses likely infect, the team determined that about 1,200 of the viruses might be involved in carbon export — the process by which carbon gets extracted from the atmosphere, incorporated into marine organisms and then "exported" to the deep sea as those organisms sink to the seafloor after death.
The deeper these carbon stores sink, the longer they're likely to remain stored in the ocean before being cycled back into the atmosphere, according to the Monterey Bay Aquarium Research Institute. For this reason, carbon export is an important factor that scientists incorporate into models of climate change. The new study suggests that the infection of marine organisms by RNA viruses may be another, previously unacknowledged factor driving carbon flux in the oceans, in that the viruses alter the cellular activity of the hosts they infect.
RNA viruses may also drive carbon flux by splitting their hosts open and spilling sequestered carbon into the ocean, Wilhelm said, since viruses often burst out of their hosts after rapidly replicating inside them.
Originally published on Live Science.

Nicoletta Lanese is the health channel editor at Live Science and was previously a news editor and staff writer at the site. She is a recipient of the 2026 AHCJ International Health Study Fellowship, with a project focused on antibiotic stewardship practices in Japan and the U.S. They hold a graduate certificate in science communication from UC Santa Cruz and degrees in neuroscience and dance from the University of Florida. Beyond Live Science, Lanese's work has appeared in The Scientist, Science News, the Mercury News, Mongabay and Stanford Medicine Magazine, among other outlets. Based in NYC, she also remains involved in dance and performs in local choreographers' work.