Dark regions of the genome may drive the evolution of new species
The findings suggest a way to rescue "doomed" animal hybrids.

Genetic "dark matter" may drive the emergence of new species, new research finds.
These long, repeating stretches of the genome, called satellite DNA, may ultimately prevent incompatible animals from mating by scrambling the chromosomes in their hybrid babies, according to the study. And if animals from different populations can't mate, they will diverge over time, leading to speciation.
Just 1% of the 3 billion letters, or nucleotides, in the human genome make the proteins that determine traits such as eye color and height. Other stretches of DNA may tell the body how many copies of a protein to make, or turn genes on or off in different tissues, among other functions. Yet nearly 10% of the human genome is composed of long, repeating stretches of satellite DNA that, for many years, scientists didn't think did much of anything, said study co-author Madhav Jagannathan, currently an assistant professor at the ETH Zurich Institute of Biochemistry in Switzerland.
Related: Genes of 500-million-year-old sea monsters live inside us
"Satellite DNA repeats were very abundant in species and widely observed in eukaryotes," or life-forms with cell nuclei, Jagannathan told Live Science in an email. "Despite this, they were largely dismissed as junk DNA."
However, in a 2018 study, Jagannathan, who was then at the Massachusetts Institute of Technology (MIT), and his former postdoctoral adviser, biologist Yukiko Yamashita, also at MIT, discovered that some of this DNA served a critical purpose: It organizes DNA within a cell's nucleus. That study found that certain proteins grab DNA molecules and arrange them in densely packed bundles of chromosomes called chromocenters. Satellite DNA, they found, tells these grabby proteins how to bundle and organize chromosomes.
In the newest study, published July 24 in the journal Molecular Biology and Evolution, Jagannathan and Yamashita found another role for satellite DNA: driving speciation. The team was investigating fertility in the fruit-fly species Drosophila melanogaster. When the researchers deleted a gene that codes for a protein called prod, which binds to satellite DNA to form chromocenters, the flies' chromosomes scattered outside the nucleus. Without the ability to correctly organize chromosomes, the flies died.
Get the world’s most fascinating discoveries delivered straight to your inbox.
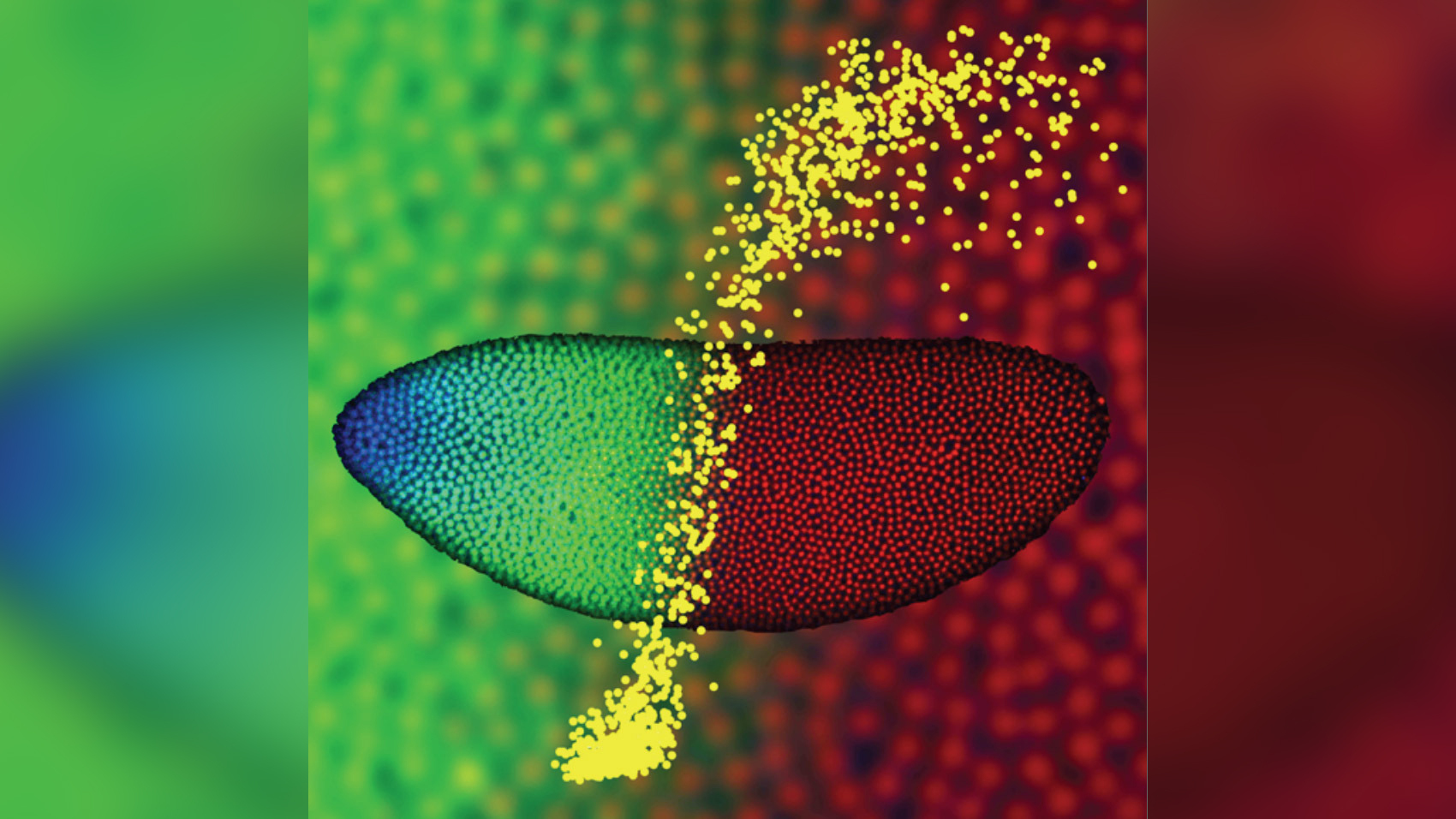
A 2-hour-old fly embryo, with DNA shown in red, a signaling protein called Bicoid in blue and another protein (Hunchback) shown in green, which play a key role in the differentiation of the head and thorax. New research shows that satellite DNA may play a role in such fruit fly embryos developing normally; flies of different species with incompatible satellite DNA-binding proteins will have cells with misshapen nuclei and chromosomes scattered throughout the cell.
This was fascinating, Jagannathan said, because the deleted protein is unique to D. melanogaster. That meant that these rapidly evolving satellite DNA sequences must also have rapidly evolving proteins that bind to them.
To test this idea, Jagannathan bred D. melanogaster females with males of a different species, Drosophila simulans. As expected, the hybrids did not live long. When the researchers looked into the flies' cells, they saw misshapen nuclei with DNA scattered throughout the cells, just as they had when they deleted the prod protein in previous experiments.
So why does that mean satellite DNA could drive speciation? The team suspects that, if satellite DNA evolves quickly and two creatures make different satellite-DNA-binding proteins, they won't produce healthy offspring. As chromocenter binding proteins and satellite DNA segments evolve differently in separate populations or species, this incompatibility could arise rather quickly.
To test this hypothesis, they mutated satellite DNA-binding genes that led to the incompatibility in both parents. When they rewrote the flies' genomes to be compatible, they produced healthy hybrids.
RELATED CONTENT
Such satellite DNA disagreements could be a big factor in the evolution of new species, Jagannathan suspects. He hopes further research can test their model of hybrid incompatibility with other species. Ultimately, this research could lead to a way for scientists to rescue "doomed" hybrids, or hybrids that don't survive long after birth. This could pave the way for using hybridization as a method for rescuing critically endangered species, such as the Northern White Rhino, of which only two females survive.
Ultimately, the new research confirmed Jagannathan's hunch that satellite DNA served a purpose.
"I thought that there was no way evolution could be so wasteful," Jagannathan said.
Originally published on Live Science.

Cameron Duke is a contributing writer for Live Science who mainly covers life sciences. He also writes for New Scientist as well as MinuteEarth and Discovery's Curiosity Daily Podcast. He holds a master's degree in animal behavior from Western Carolina University and is an adjunct instructor at the University of Northern Colorado, teaching biology.

 Live Science Plus
Live Science Plus




