Learning from Earth's Smallest Ecosystems (Kavli Hangout)
Get the world’s most fascinating discoveries delivered straight to your inbox.
You are now subscribed
Your newsletter sign-up was successful
Want to add more newsletters?
Join the club
Get full access to premium articles, exclusive features and a growing list of member rewards.
Alan Brown, writer and blogger for the Kavli Foundation, contributed this article to Live Science's Expert Voices: Op-Ed & Insights.
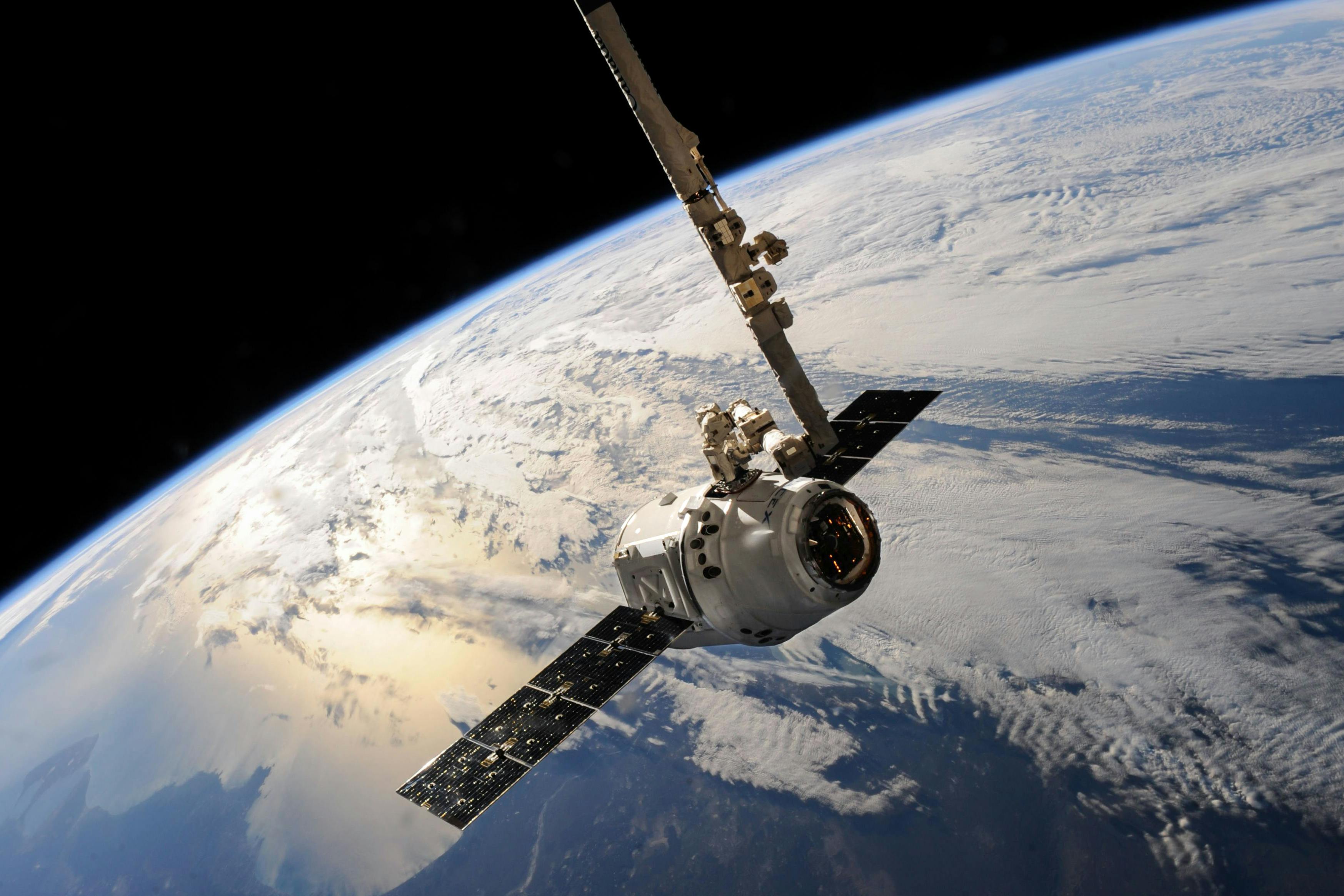
From inside our bodies to under the ocean floor, microbiomes — communities of bacteria and other one-celled organisms — thrive everywhere in nature. Emerging at least 3.8 billion years ago, they molded our planet and created its oxygen-rich atmosphere. Without them, life on Earth could not exist.
Yet we know surprisingly little about the inner workings of nature's smallest and most complex ecosystems.
Article continues belowMicrobiomes have a great deal to teach us. By learning how members of microbiomes interact with one another, scientists might discover innovative green chemistry and life-saving pharmaceuticals, or learn how to reduce hospital infections, fight autoimmune diseases, and grow crops without fertilizers or pesticides.
The sheer complexity of microbiomes makes them difficult to study by conventional biochemical means. Nanoscience provides a different and complementary set of tools that promises to open a window into this hidden world. [The Nanotech View of the Microbiome]
Earlier this month, The Kavli Foundation hosted a Google Hangout with two leaders in the emerging applications of nanoscience for studying microbiomes. They discussed the potential of natural biomes, why they are so difficult to understand, and how nanoscience may help us unlock microbiome secrets.
Joining the conversation were:
Get the world’s most fascinating discoveries delivered straight to your inbox.
Eoin Brodie, a staff scientist in the Ecology Department at Lawrence Berkeley National Laboratory. He was part of the team that pioneered a device capable of identifying thousands of the bacterial species found in microbiomes, and is currently developing ways to combine data from many different types of measurement tools into a more coherent picture of those ecosystems.
Jack Gilbert is a principal investigator in the Biosciences Division of Argonne National Laboratory and an associate professor of ecology and evolution at the University of Chicago. He has studied the microbiomes of hospitals and is working on ways to use nanostructures containing bacteria to help infants fight immune diseases.
Below is a modified transcript of their discussion. Edits and changes have been made by the participants to clarify spoken comments recorded during the live webcast. To view and listen to the discussion with unmodified remarks, you can watch the original video.
The Kavli Foundation: So let's start with an obvious question, what exactly is a microbiome?
Eoin Brodie: A microbiome is a connection of organisms within an ecosystem. You can think of the ecosystem of microbes in the same way you think of a terrestrial ecosystem, like a tropical forest, a grassland, or something like that. It is a connection of organisms working together to maintain the function of a system.
Jack Gilbert: Yes. In a microbiome, the bacteria, the archaea (one-celled organisms similar to bacteria), the viruses, the fungi, and other single-celled organisms come together as a community, just like a population of humans in a city. These different organisms and species all play different roles. Together, they create an emergent property, something that the whole community does together to facilitate a reaction or a response in an environment.
TKF: How complex can these microbiomes? Are they like tropical forests? Are they more complex, less complex?
J.G.: The diversity of eukaryotic life — all the living animals and plants that you can see — pales into insignificance beside the diversity of microbial life. These bacteria, these archaea, these viruses — they've been on the earth for 3.8 billion years. They are so pervasive, they have colonized every single niche on the planet.
They shaped this planet. The reason we have oxygen in the atmosphere is because of microbes. Before they started photosynthesizing light into biomass, the atmosphere was mostly carbon dioxide. The reason the plants and animals exist on Earth is because of bacteria. The diversity of all the plants and animals — everything that's alive today that you can see with your eyes — that's a drop in the proverbial ocean of diversity contained in the bacterial and microbial world. [Can Microbes in the Gut Influence the Brain?]
E.B.: We tend to think of the earth as being a human planet and that we're the primary organism, or the alpha species. But we're really passengers, we're just blow-in's on a microbial planet. We're recent, recent additions.
TKF: You both wax so poetic about it. Yet we know so little about microbiomes. Why is it so hard to understand what goes on in these ecosystems?
E.B.: Jack eluded to it. The first problem is that microbiomes are very small. We can't see them, and it's very difficult to understand how things work when you can't see them. So tools are needed to be able to see these organisms.
We also can't grow them. It's very hard to bring them from the natural ecosystem into the lab for study. Probably less than one percent, depending on the ecosystem, can actually be cultivated on growth media in the lab so that we can do experiments and understand what functions they carry out. That leaves 99 percent — the vast majority of the microbes on Earth and most of their ecosystems — unknown to us, apart from their DNA signatures and things like that.
Now, Jack has pioneered DNA analyses. When you look at the DNA signatures from these environments, there are all these new organisms, new proteins, and new functions that we have never really seen before. This has been called earth's microbial dark matter. Just like dark matter and energy in the universe, this has been unknown to us, but it is extremely important if the planet — and humans — are to continue to function.
TKF: So, what makes it so hard to grow these microbes in a Petri dish?
E.B.: They're very fussy. You can think of it that way. They don't like to eat the food that we give them, in many cases. They eat things that we don't know they can eat. They breathe things that we don't know that they can breathe.
We breathe oxygen, they breathe oxygen, but they also breathe nitrates, iron, sulfur, even carbon dioxide. Getting the right concentrations and combinations of what they eat and breathe is very difficult.
In some cases, even if you can work that out, there may be something that they need to get from another member of the ecosystem. That member may supply an essential nutrient or a cofactor for them to grow.
So getting all of those possible permutations and combinations right is extremely challenging. A lot of people are working on it, and there's a lot of expertise being put into this, but it's extremely difficult and complicated.
J.G.:& That's an interesting point. I liken it to having a baker. You know, if you have a baker in a human community, the baker needs somebody who can make the flour, somebody who can provide a bit of yeast, and someone who will buy the bread. They exist as a network of individuals living in a community.
If you take the baker out of the community, he or she cannot make the bread and so they are no longer a baker. Removing a microbe from its community reduces the likelihood that it will be able to perform the roles and tasks that it does in that environment.
So it's almost like you don't want to try and grow these things in isolation. Because, while isolating them makes our job as a microbiologist easier, it's also much more difficult to understand what they actually do in the environments in which they live. We can't figure that out in isolation because they are community players.
TKF: What are some of the tools that we can use today to look at microbiomes? Is there a state of the art?
J.G.: So I'll take on that. I mean this is a very dynamic evolving field. It is not a field where everyone seems to rest on their laurels.
To understand microbes, we have a couple of tools that are available to us. One of those tools is genomics, so we can sequence the genome of bacteria, archaea, viruses and fungi, just as we've done for the human genome.
The second one is the transcriptome, which looks at RNA, a transient molecule that creates the cell by translating what's in the genome into proteins. That's useful, because it tells us which genes are being turned on and off when we put those microbes under different conditions.
Then we have the proteome, the proteins that actually make up the cell. They are the enzymes that enable the organism to interact with its environment, to consume its food, to respire carbon dioxide, oxygen or iron, and so on.
Then you have the metabolome, the metabolic molecules living organisms consume as food and produce as waste products.
The genome, transcriptome, proteome, and metabolome are four of the tools in our toolbox that we can actually use to examine the microbial world. But they are by no means the limit of our tools or our goals. We have ambitions far beyond just examining those components. Eoin is developing some of these, and maybe Eoin, you want to jump in now?
E.B.: Yes, I'd add to that. The challenge of understanding the microbiome, and even individual microbes, is that they're just so small. They're complicated and small, so understanding their activity — their transcriptomes or proteins or metabolites — at the scale at which they exist, is extremely challenging.
All the technologies that Jack mentioned are being developed with larger organisms in mind. Scaling them down to deal with the size of microbes, but then increasing their throughput to deal with the complexity of microbes, is a huge, huge challenge.
I'll give you an example. When you look at the activity of an ecosystem, say a tropical forest, you look at the distribution of trees and animals, and look for the association between the vegetation and animals.
So if you want to understand insects, you have a space in mind. You think, "This lives near this. It interacts in this area." So there's an interaction, a fundamental association between those members of the ecosystem.
The way we typically looked at microbiomes — though this is changing now — was to mash up the entire forest in a blender. Then we would sequence all of the DNA, and look at the RNA and proteins, and the metabolites.
Then we try to go back and say, "This tree is interacting with this insect." Whereas, in reality, that tree is hundreds or thousands of kilometers away from that insect, and they never see each other.
That's the problem we have in the microbiome. When we mash up those organisms to look at their DNA, RNA, proteins and metabolites, we get rid of that spatial structure and its associations. And we lose the importance of space in terms of facilitating interactions. [The Nanotech View of the Microbiome (Kavli Roundtable)]
So, really, I think the next wave in microbiome research has to target this microbial activity and interactions at the scale of the microbe. Do they see each other? Do they interact, and how do they interact? What chemicals do they exchange, and under what conditions? I think that's the real challenge. That's why we're talking to the Kavli Foundation, because that's where nanoscience comes in.
TKF: This is an excellent transition to my next question: How do we use nanoscience to learn about microbiomes? For example, could we use some of the same nanoscale probes we are developing to study the brain to, say, investigate microbiomes in the ocean or soil?
E.B.: I think there are some interesting parallels. I mean, you can think of the brain as this extremely complicated network of neurons. The BRAIN Initiative is attempting to map those neurons and to follow their activity.
Similarly, the microbiome is a network of interacting organisms that turn on and turn off. The connections and the structure of that network are extremely important to the functioning of the system, just as it is for the functioning of the brain.
For the BRAIN Initiative, people got together and said, "Well what do we need to do to look at electrical charge and electrical flow through neurons, noninvasively, and in real time?" And they came up with some technologies, that can potentially, do remote sensing on a very small scale, and watch how the system changes noninvasively.
So, one approach to understanding the brain is to use external imaging, and another approach is to embed sensors.
In the BRAIN Initiative some sensors are being developed here at Berkeley lab and elsewhere that use RFID — radio frequency identity — technology. They are similar to tags used to track shipping containers, goods in department stores, and things like that. They both transmit information and harvest energy from radio frequencies, so they're autonomous devices. I think that the challenge now is coupling that technology to sensors that can monitor something in the environment and send that information autonomously — no batteries required — to receivers. Then, if these sensors are distributed in an intelligent way, just like with GPS, you can triangulate where that information is coming from.
How could you use this to understand a microbiome? Well, the sensors that are being developed are still relatively large scale, about one square millimeter in size. That's pretty small for us, but very large for a microbe.
So you can think about this in soil. Let's say we want to understand what happens when a root grows through soil. The root stimulates microbes, and there are ten times more microbes near the root than there are away from the root in soil. They all have different chemistries and different functions that are very important for the nutrition and health of the plant.
If you could distribute very small sensors in the soil and have them sense things like carbon from roots or oxygen consumed by microbes, then you can build a three dimensional picture of how the soil microbiome is changed and altered as a root moves through the soil. That's one example of how advances in other fields, driven by nanotechnology, could be applied to microbiome.
TKF: These RFID sensors would be based on semiconductor chips, right? So you could take a wafer, make a lot of them cheaply, distribute them in the soil, and get a picture you couldn't get any other way?
E.B.: Yes. There's an emerging field called predictive agriculture. It's like personalized agriculture, where fertilizer addition, for example, in a field would not be uniform. Instead, you would deliver the fertilizer where it's needed. You would irrigate the field exactly where it's needed. So you have this massive network of distributed autonomous sensors, and that would allow us to more efficiently use fertilizer. Then it wouldn't be leached or lost from the system, and cause water pollution and things like that. These examples are not on a microbial scale, but microbial processes control the availability and uptake of these fertilizers.
TKF: Thank you. Hold that thought and we'll come back to it in a few moments. In the meantime, Jack has been studying microbiomes in a new hospital to see how they evolve and affect the spread of disease. Could you tell us what you are doing, and how nanotechnology might help?
J.G.: Yes. The microbes that exist in a hospital have been a focus of clinicians and medical researchers for a couple of hundred years. Ever since we uncovered that bacteria might actually be causing disease, we've been trying to eradicate as much microbial life as possible.
That paradigm is shifting to one where we're more interested in trying to understand how bacterial communities in a hospital may facilitate the spread of disease and antibiotic resistance, and maybe promote health as well.
We've been going into hospitals and, with a very, very high temporal resolution, exploring how their bacterial communities change over time. So, looking at a scale of hours to days, we're trying to understand how — when a patient moves into a new room to have an operation or to undergo a procedure — the microbes that are already in that room affect the outcome of the patient's stay in the hospital. We want to know if it makes them either healthier or sicker.
So, we've been cataloging the microbes at these very fine scales. And what we see is an exchange between the bacteria in the room and inside the patient's body.
But we've also discovered that the vast majority of bacteria that we would normally associate with so-called healthcare-associated infections — pathogens that we thought people acquire during hospital stays — appear to be bacteria that patients brought into the hospital themselves. They're bacteria that we have inside us.
Remember, we have one hundred trillion bacteria living inside us. They weigh about two pounds, about the same as the brain. So if you think that the BRAIN Initiative is important, well maybe a microbiome initiative would also be important, because it weighs about the same as the brain.
The human microbiome has a lot of players. Most of them are friendly to us, but they can turn on us too. I liken this to a riot spreading in the city. You know, if you take things away from people, they will generally rise up and try to overthrow the very thing which was supporting them in the first place.
Microbes are the same way. We give a hospital patient antibiotics and radiation therapy to kill bacteria. Then we cut open his or her intestine and expose the bacteria to oxygen, which they don't like, and stitch the gut back up. When we look at the bacteria, we see that previously friendly bacteria have started to riot. They've been insulted so many times by the patient's treatment that they've decided that they've had enough. Then they go and attack the host to regain the resources which are being taken away from them.
This is very important. Understanding a patient's hospital stay from the microbes' perspective is helping us to design better ways to treat patients and reduce the likelihood that those microbes inside us will rebel, attack us, and make us sick.
Nanotechnology is helping us to achieve a finer scale of visual resolution, so we can see exactly when, during a surgical procedure, bacteria go rogue and start to attack the host, and the molecular mechanisms that underpin that behavior.
We have a great example that we found by placing nanoscale molecular biosensors in the gut. It measures phosphate levels. Phosphate is a very important molecule that is used to create the DNA and proteins in our body, and in the cells of those bacteria.
When the phosphate level drops below a certain threshold, the microbes turn on a mechanism to acquire phosphate from their environment. And where's the best source of phosphate? It's in the gut lining of their host. So they migrate to the gut and start to break down the human cells. We experience that as a several pathogenic infection, which often kills us.
Because we understand that process, we are developing mechanisms to release phosphate at exactly the right time during surgery to prevent those bacteria from ever experiencing that phosphate reduction. To do those micro phosphate releases, we're developing nanotech scaffolds to hold phosphate, and placing them into the gut during surgery. This will reduce the likelihood that microbes will become pathogenic.
TKF: Not only is that interesting, but it leads one of our viewers to ask whether we can adjust microbiomes so that they can target diseases and other human conditions. Can they go beyond just adjusting acidity or phosphate levels and do something more aggressive?
J.G.: Yes. The case where we've had the best success is in treating chronic infections caused by Clostridium difficile bacteria. C. diff infections are chronic gastrointestinal infections. Our treatments use a shotgun approach. We take the bacteria from a healthy person and transplant them into somebody with a chronic C. diff infection. That's overridden the C. diff infection, and established a healthy microbiome in the patient's gut so that he or she is no longer sick.
The Chinese did this about 2,000 to 3,000 years ago. They called it yellow soup, and they fed the stool from a healthy person to a sick person, and that made the sick person healthy. We just rediscovered this process, and we are now applying it in a more clinical setting.
So far, it's a very untargeted approach. What we're trying to do with our research arm, American Guts, and programs associated with autism, Alzheimer's, and Parkinson's, is to identify specific bacterial community members that are either absent or overgrown in those patients. Then we want to explore how to adjust them — maybe we implant one that is missing or knock one back that is over-grown, to make that person healthier.
E.B.: I'd like to add something to that. There's an interesting analogy, I think, in what we're doing for C. diff — fecal transplants — and restoration ecology. That's where you weed out an invasive plant species and plant another species to out-compete that invasive plant species. It's the exact same process, so the same ecological principles and ecological theory that's used in restoration ecology can be used in medicine. In some cases, it may not be as simple as removing one organism or adding one or two other organisms. It might be a community function, where we may actually need that complexity to be able to out-compete the organism that's causing the disease.
J.G.: That's a really interesting point. Both Eoin and I are microbial ecologist at our core. I started out in marine microbial ecology, and now I work in soils, plants, humans, and disease. Eoin does the same. And both of us can apply the ecological principles of microbes to any environment because microbes are everywhere.
TKF: Good. So, Eoin, we have two questions for you from our audience. The first involves agriculture. A viewer want to know whether nanoscience help us alter microbiomes in ways that change how we grow, fertilize, and protect plants from pests?
E.B.: That's a great question, and I think a really timely one as well. The world population is seven billion, heading to nine, and then 11 billion. We're going to run out of fertilizer, we're going to run out of space to grow food, and we're running out of water — we're in a severe drought in California. These are our challenges, feeding a global population and providing fuel for a global population.
The things microbes and nanotechnology can do mainly revolve around improving the resistance of plants to stresses, such as drought. Microbes can help plants acquire water. For example, mycorrhiza fungi can increase the root system, improve its drought tolerance, and improve nutrition.
We can also identify bacteria that can produce fertilizer in or near the plant. So bacteria that can take nitrogen from the atmosphere and fix nitrogen can potentially offset the use of nitrogen fertilizer, which takes a lot of energy and causes a lot of pollution to manufacture.
Bacteria can also mine critical minerals from the soil. We can have bacteria growing with the plants that acquire phosphorous, like Jack was saying. We can choose bacteria so that they mine more phosphorous than they need and supply that to the plant.
All of these things would reduce our reliance on mining phosphorous from strip mines or using five percent of our world's energy to product nitrogen fertilizer. I think it's a big, big challenge.
Nanotechnology, as I mentioned earlier, can be used to characterize these organisms and understand how they work. We can also build sensor systems to identify when nutrients are limiting growth. So instead of spreading nutrients and fertilizer in a very inefficient way, we can use it in a very targeted, specific, and much more sustainable way.
TKF: Can we take a step beyond that, and perhaps use microbiomes to control pests?
E.B.: Actually, that's been done for a long time. As you know, there are GMO crops out there that have taken genes from microbes that are used to kill insects. This could be carried out in a more natural way, as well, for example, by growing these bacteria with the plants and potentially inhibiting insects from grazing and feeding on the plants. We can learn a lot from nature. Nature has already developed these strategies for pest control, and we can learn from that to design our protections in a more, controllable and intelligent way.
TKF: Another question from a viewer: Is it possible to make an artificial microbiome community do a particular task?
J.G.: Yes. We've actually been working in that area, trying to create what we call a simple minimal community. This is a community of organisms that performs a task, such as creating acetate or generating hydrogen or butanol as potential biofuel source. So we're looking at microbes that grow on the surface of cathodes, and take raw electrons from those cathodes and integrate them with a carbon dioxide source, such as blue gas from a factory. We want to create a community that drives it's metabolism towards a set goal.
That will take a mathematical modeling approach. So metabolic modeling, trying to synthesize in a computer how these microbes interact to release a certain product. So, in that sense, you need nanotechnology to sense the metabolic relationships that exist between those organisms, so that you can engineer that community towards producing a particular product. That's going to be very important to achieve biotechnology results.
E.B.: Actually, I've got to turn that question on its head. I would like to take a natural microbial community and stop it doing something, in certain cases.
Let's say, for example, you've got cattle livestock. They are a significant source of global methane that contributes to global warming. Part of that is because of their diets, which provide an excess energy. That results in increased hydrogen, which results in a lot of methane, and cows release a lot of methane.
So, could we go in and use targeted synthetic biology or chemical interference approaches to stop the production of methane? To alter the balance of the cow's rumen, the cow's gut microbial ecosystem? We could not only inhibit methane production, but improve nutrition to the animal, because it's microbes that control the flow of energy to the animal from the food that it eats.
It's a complicated ecosystem, but specifically tweaking it for the benefit of the animal and the benefit of the planet, is an interesting challenge and there are people working on that.
J.G.: I'd like to take that exact system and apply it to coal, in order to make more methane that we can then capture and pump into people's homes as biofuel.
TKF: Interesting thought. I have another question from a viewer, and Jack, I think you are the one to answer this. She has of experimental treatments that involve implanting health gut bacteria into people with autism. Why might this work? And will this be something that we see soon?
J.G.: The bacteria in our gut have an impact upon neurological behavior — the way we behave — through our immune system. They elicit a certain immune response in our gut, which feeds back on our nervous system to create a certain characteristic behavior in our brain.
We've known this in animal models for a number of years now. We're just starting to understand the extent to which neurological diseases, such as autism, Parkinson's, and conditions such as Alzheimer's, are attributable to a disruption in the bacterial community in somebody's intestine.
There have been several experiments with very low numbers of children. In several cases in South America and a number in Australia, the children have had a fecal microbiome transplant, a healthy microbial community implanted into their own gut.
The results are variable, and not exactly something that you would want to try at home. But they do hint, in some instances, of a favorable outcome where the child's neurological disorder is lessened, or significantly reduced.
There are groups at Cal Tech are generating probiotics, particular bacteria species, that they hope to add to a child's diet or put into a capsule that can be swallowed. They seem to have a benefit in reducing the neurological abnormalities associated with autism, though they are still in their early days.
TKF: That leads to another question I wanted to ask you. Jack, you're also working on encapsulating microbiomes in some sort of nanostructure and applying them to homes or offices. Your hope is that these biomes will expose people to microbiomes that will help their immune system develop resistance to these neurological problems. Could you tell us about that?
J.G.: Yes, we're working on animal models at the moment. Imagine recreating structures that these animals can interact with. Imagine I build you a building that was biologically alive, where the walls were deliberately teeming with a healthy microbial community.
Now, we have only a very limited idea what healthy means, but essentially what we're doing is creating structures, 3D printable structures, impregnated with certain nutrients. We're working with Ramille Shah at Northwestern University to create a 3D structure which allows that bacterial community to thrive.
We can then introduce these structures into a mouse's cage. The bacteria associated with the 3D surface will colonize that mouse, and reduce certain abnormalities that we see in that mouse, such as an allergy response. So we've been growing bacteria which can produce a chemical that, once released into the gut of the mouse, will form a colony and reduce the likelihood of that mouse having a food allergy.
I'm also working with Cathy Nagler at the University of Chicago. We're hoping to prove that we don't have to pump kids full of probiotics. Instead, we can just redesign homes, schools, and maybe daycare centers, so that children will get an appropriate microbial exposure that would mirror how they would have grown up if they were in a natural ecosystem. Hopefully, that will be the future of architecture.
E.B.: And, you know, as a possible alternative, we can send our kids outside to play more.
J.G.: You got it.
E.B.: Not bad.
Follow all of the Expert Voices issues and debates — and become part of the discussion — on Facebook, Twitter and Google+. The views expressed are those of the author and do not necessarily reflect the views of the publisher. This version of the article was originally published on Live Science.
 Live Science Plus
Live Science Plus














