Protective Shell of a Virus Imaged

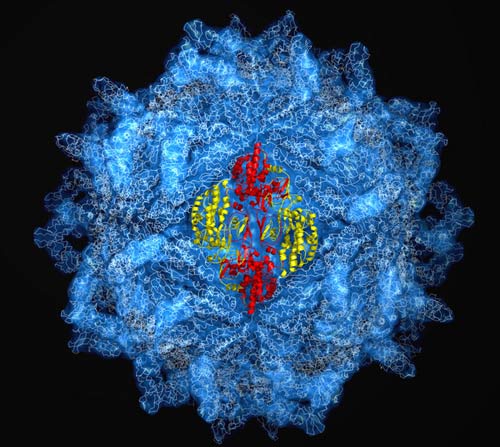
Scientists have painted the clearest picture yet of the coat of proteins that surround hundreds of known viruses with a new image that pinpoints the locations of some 5 million atoms that make up the protective shell.
The image, created by high energy X-rays and detailed in the Feb. 16 issue of the journal Proceedings of the National Academy of Sciences, could help researchers find better ways to fight viral infections.
The shell of proteins, called the "capsid," encases the genome of the viruses. While we humans, and most other living creatures, have genomes composed of DNA, the genomes of these viruses consist of DNA's cousin, RNA (ribonucleic acid). RNA is much like DNA except that it has a slightly different set of nucleic acids (the chemical "letters" that spell out our genetic code).
Another group of researchers recently decoded the genome for the common cold.
Viruses' capsids come into play because viruses can reproduce themselves only by invading a host cell and hijacking its biochemical machinery. But when they invade, viruses need to seal off their genetic payload to prevent it from being destroyed by the cell's protective mechanisms.
"When these viruses invade cells, the capsids get taken inside and never completely break apart," said lead researcher Jane Tao, of Rice University in Houston.
{{ video="090109_VirusMotor" title="Special Delivery: Antibiotic Viruses Could Kill Bacteria" caption="Researchers discover powerful molecular-scale motors that drive viruses." }}
Get the world’s most fascinating discoveries delivered straight to your inbox.
Though there are more than 5,000 known viruses, including whole families that are marked by wide variations in genetic payload and other characteristics, most of them use either a helical or a spherical capsid.
In their attempt to map precisely the spherical variety, Tao and colleague Junhua Pan, a postdoctoral researcher at Rice, first had to create a crystalline form of the capsid that could be X-rayed. They chose the much-studied Penicillium stoloniferum virus F, or PsV-F, a virus that infects the fungus that makes penicillin.
Though PsV-F does not infect humans, it is similar to a rotavirus and others that do.
"Spherical viruses like this have symmetry like a soccer ball or geodesic dome," Pan said. "The whole capsid contains exactly 120 copies of a single protein."
Previous studies had shown that spherical capsids contain dozens of copies of the capsid protein, or CP, in an interlocking arrangement. The new research identified the sphere's basic building block, a four-piece arrangement of CP molecules called a tetramer.
By deciphering both the arrangement and the basic building block, the research team hopes to learn more about the capsid-forming process.
"Because many viruses use this type of capsid, understanding how it forms could lead to new approaches for antiviral therapies," Tao said. "It could also aid researchers who are trying to create designer viruses and other tools that can deliver therapeutic genes into cells."
The research was supported by the National Institutes of Health, the USDA, the Welch Foundation, the Kresge Science Initiative Endowment Fund, the Agouron Foundation and the San Diego Supercomputer Center.
- How Viruses Invade Us
- How Viruses Work: Natural Motors Revealed
- Virus News and Information


 Live Science Plus
Live Science Plus





