
Network of Mics Helps Track Sea Creatures on the Move

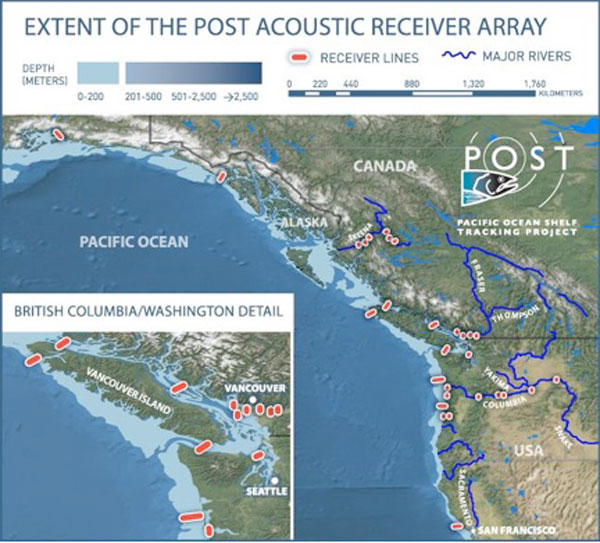
Microphones deployed along hundreds of miles of coastline are now tracking thousands of marine animals during their journeys through rivers and in the ocean.
For the past eight years, scientists have been tagging nearly 16,000 animals with acoustic transmitters that each broadcast a unique identifier. At the same time, researchers have deployed more than 400 acoustic receivers in lines situated at strategic locations along the West Coast of the United States, running from the shore and out to the edge of the continental shelf.
The resulting array, dubbed the Pacific Ocean Shelf Tracking (POST) network, stretches more than 1,800 miles (3,000 kilometers), extending from Alaska through British Columbia to California. When a tagged creature crosses a line in this network, it gets detected by at least one acoustic receiver on that line, data which researchers can later download wirelessly.
In this fashion, scientists can track where and when the creatures go over time. And that's not all: Since the lines are laid out in such a way that nearly all tagged animals of a group are detected as they pass a line, researchers can even estimate whether any members might have died.
"People have very limited interactions with and understanding of animals under the water it's a completely different environment, and we're limited in our ability to get down there and see what life is doing," said Jonathan Thar, research program coordinator for the POST Project. "Technology like this really captures the imagination for me when it comes to what we might learn about a whole new world we don't know that much about."
The tags range in size from a tube of lipstick to an almond. The smallest devices can be used to tag creatures just 5 inches (12.5 centimeters) long, while bigger tags with larger batteries can track creatures for up to seven years at a time. They can be detected from a range of 1,300 feet (400 meters) or more, much farther than typical radio tags, which have only a range of a few yards at most. While satellite tags can be used to track creatures around the world, they cost $5,000 to $10,000 each, as opposed to $300 to $350 each for the acoustic tags.
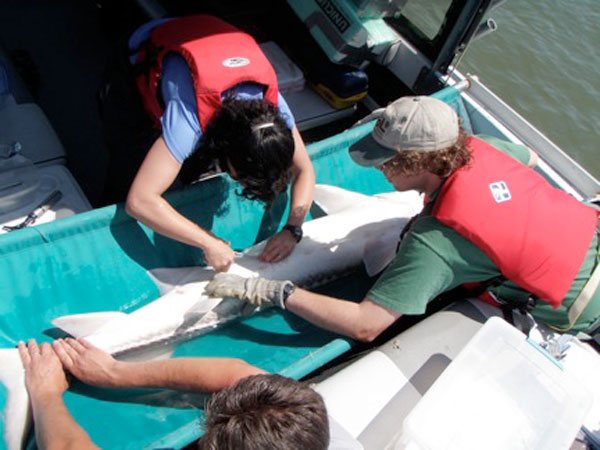

"Sometimes tracking the animals brings surprising results that turn conventional wisdom upside down," said Jim Bolger, POST's executive director.
Get the world’s most fascinating discoveries delivered straight to your inbox.
For instance, researchers using the array have unexpectedly discovered that juvenile Pacific salmon fared just as well negotiating the heavily dammed Columbia River as they did going down the free-flowing Fraser River, and even survived better in the dammed Chinook River than on the Fraser. This challenges the widely held assumption that dams pose the biggest threat to salmon recovery it remains uncertain whether the Fraser has some problem that kills salmon as much as heavy dams might, or if there are factors other than dams playing a larger, unsuspected role in the life and death of salmon.
Scientists have also revealed that rockfish and lingcod took a liking to Alaska's first intentional artificial reef near Whittier, Alaska, in northwestern Prince William Sound. Fish released at the reef continued to stay there as often as they tried returning to their original homes in natural reefs nearby. This suggests artificial reefs could help serve as viable dwellings for these creatures in case their normal habitats become polluted or otherwise too dangerous for them.
The large-scale network is available free of charge, and its database, complete with tools for mapping and visualizing data, serves as a clearinghouse for knowledge gathered across the array and similar local networks. Anyone, not just scientists, who is curious about what marine life is doing nearby can monitor the creatures as well.
The project's long-term goal is to operate a series of lines of acoustic receivers from the Baja Peninsula to the Bering Sea. In terms of what's next for tagging, "the sky's the limit," Thar said. "People are now figuring out ways to tag lion's mane jellyfish, for instance."
The POST Project detailed its findings online Aug. 31 in the online journals PLoS ONE and PLoS Biology.


 Live Science Plus
Live Science Plus





