Largest human family tree ever created retraces the history of our species
The tree is based on thousands of human genome sequences.

A new, enormous family tree for all of humanity attempts to summarize how all humans alive today relate both to one another and to our ancient ancestors.
To build this family tree, or genealogy, researchers sifted through thousands of genome sequences collected from both modern and ancient humans, as well as ancient human relatives, according to a new study published Thursday (Feb. 24) in the journal Science. These genomes came from 215 populations scattered across the world. Using a computer algorithm, the team revealed distinct patterns of genetic variation within these sequences, highlighting where they matched and where they differed. Based on these patterns, the researchers drew theoretical lines of descent between the genomes and got an idea as to which gene variants, or alleles, the common ancestors of these people likely carried.
In addition to mapping out these genealogical relationships, the team approximated where in the world the common ancestors of the sequenced individuals lived. They estimated these locations based on the ages of the sampled genomes and the location where each genome was sampled.
Related: In photos: A nearly complete human ancestor skull
"The way that we've estimated where ancestors live is, in particular, very preliminary," said first author Anthony Wilder Wohns, who was a doctoral student at the University of Oxford's Big Data Institute at the time of the study. Despite its limitations, the data still captured major events in human evolutionary history. For example, "we definitely see overwhelming evidence of the out-of-Africa event," meaning the initial dispersal of Homo sapiens from East Africa into Eurasia and beyond, said Wohns, who is now a postdoctoral researcher at the Broad Institute of MIT and Harvard.
The method the researchers used "works well to refine known ancestral locations and, as sampling improves, it has the potential to identify currently unknown human movements," Aida Andrés, an associate professor in the Genetics, Evolution and Environment Department at the University College London (UCL) Genetics Institute, and Jasmin Rees, a doctoral candidate at the UCL Genetics Institute, wrote in a commentary, also published in the journal Science on Thursday. So, in the future, when more data become available, such analyses could potentially reveal chapters of human history that are currently unknown to us.
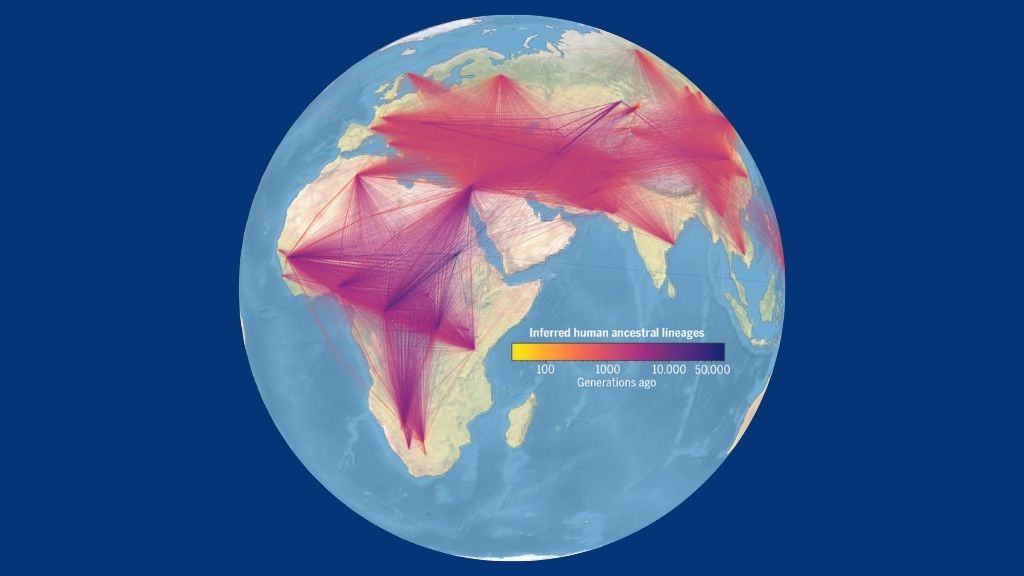
Each line in this figure represents an ancestor-descendant relationship in the new genealogy of modern and ancient genomes. The width of a line corresponds to how many times the relationship was observed in the data, and the lines are colored on the basis of the estimated age of the ancestor.
Building the human family tree
To build a unified genealogy of humanity, the researchers first pooled genomic data from several large, publicly available data sets, including the 1000 Genomes Project, the Human Genome Diversity Project and the Simons Genome Diversity Project. From these data sets, they gathered about 3,600 high-quality genome sequences from modern-day humans; "high-quality" genome sequences are those with very few gaps or errors, which have been largely assembled in the correct order, according to a 2018 report in the journal Nature Biotechnology.
Get the world’s most fascinating discoveries delivered straight to your inbox.
High-quality genomes from ancient humans were harder to come by, since DNA from ancient specimens tends to be severely degraded, Wohns said. However, in digging through previously published research, the team managed to find eight high-quality ancient hominin genomes to include in their tree. These included three Neanderthal genomes, one thought to be more than 100,000 years old; a Denisovan genome roughly 74,000 to 82,000 years old; and four genomes from a nuclear family that lived in the Altai Mountains of Russia about 4,600 years ago. (Neanderthals and Denisovans are extinct relatives of Homo sapiens.)
In addition to these high-quality ancient genomes, the team identified more than 3,500 additional, lower-quality genomes with significant degradation, ranging from a few hundred to several thousand years old, Wohns said.
These degraded genomes did not factor into the main tree-building analysis, but the team sifted through the fragments to see which isolated alleles could be identified in the samples. This piecemeal data helped the researchers confirm when different alleles first cropped up in the genealogical record, since the specimens that the genomes came from had been radiocarbon dated.
Ancient genomes provide a "unique snapshot of genetic diversity in the past," which can help reveal when and where a genetic variant first appeared, and how it spread thereafter, Andrés and Rees told Live Science in a joint statement. "Whilst this study does not integrate the low-quality ancient genomes into the building of the tree, using them to inform the age of variants within the tree is still powerful for these means, and promises many exciting advances ahead."
Wohns and his colleagues used these data to double-check whether the lines of descent outlined in their family tree made sense, timing-wise — and, in most cases, they did.
Related: Unraveling the human genome: 6 molecular milestones
"It's very reassuring to see that … over 90% of the time, we are being consistent with the samples that archaeologists can radiocarbon date," Wohns said. "But there are, you know, 5[%] or 10% of these genetic variants where we see discordant estimates" as to when they first appeared, according to conflicting results from the archaeological record and the estimates made by their tree-building algorithm, he noted. In these cases, the team adjusted their tree to reflect the timing that could be confirmed through radiocarbon dating, he said.
Although it's based on just a few thousand genome samples, the team's final family tree "actually captures quite a lot about the genealogy of all of humanity," Wohns said. Using the tree as a scaffold, the team then conducted their geographical analysis, to see when and where the theoretical ancestors of their sampled populations likely lived. From this, they not only found clear evidence of the out-of-Africa migration but also uncovered potential evidence of interactions between Homo sapiens and now-extinct hominids, such as the Denisovans, he said.
For example, their results suggested that ancestors of modern humans could be found in Papua New Guinea some 280,000 years ago, hundreds of thousands of years before the earliest known evidence of modern human habitation in the region. That doesn't necessarily suggest that H. sapiens actually occupied the area that long ago, "but it does perhaps suggest that there's some genetic variation that is only found in that region, and indicates that there's a really deep ancestry there that's not found elsewhere," he said.
Some of this unique ancestry may stem from modern humans breeding with Denisovans, as was also suggested in a 2019 report in the journal Cell, which found genomic evidence of modern humans interbreeding with multiple Denisovan groups.
RELATED CONTENT
"The trees generated in this study will undoubtedly prove useful to those studying human evolution," but the methods and data used to construct said trees are "not without their limitations," Andrés and Rees wrote in their commentary. One limitation is that most genomic sequencing has been performed in Eurasian populations, so although the new study incorporated thousands of modern genomes, the data may not fully capture global genetic diversity, they told Live Science in an email. "Further integration of under-represented populations would continue to tackle this limitation," they said.
"There's a lot of uncertainty in these estimates," Wohns said of the team's recent results. "Unless we had the genome of everybody who ever lived, and where and when they lived, that's the only way that we can get the truth." The team reconstructed human history as closely as they could given the data at hand, but with more genome samples and more sophisticated software, the tree could definitely be refined, he said.
Related: Photos: Looking for extinct humans in ancient cave mud
"The nice thing about the methods we've created is that they would work with potentially millions of samples," Wohns said. "So, as we have more data, we'll get better estimates."
Wohns said he's now working to develop new machine-learning algorithms to improve the team's estimates of where and when our ancestors lived. In a separate project, he plans to employ the same tree-building method to better understand the genetic basis of human disease. He aims to do this by pinpointing the origin point of disease-related alleles and then reconstructing how and when these gene variants spread through different populations.
The same tree-building method could also be used to trace the evolutionary history of other organisms, such as honeybees or cattle, and even infectious agents, like viruses, he added.
"The power and resolution of tree-recording methods promise to help clarify the evolutionary history of humans and other species," Andrés and Rees wrote in their commentary. "It is likely that the most powerful ways to infer evolutionary history going forward will have their foundations firmly set in these methods."
Editor's note: This article was updated at 10 a.m. on Feb. 25, 2022 with additional comments from Aida Andrés and Jasmin Rees. The original article was posted at 7 a.m. EST on the same day.
Originally published on Live Science.

Nicoletta Lanese is the health channel editor at Live Science and was previously a news editor and staff writer at the site. She is a recipient of the 2026 AHCJ International Health Study Fellowship, with a project focused on antibiotic stewardship practices in Japan and the U.S. They hold a graduate certificate in science communication from UC Santa Cruz and degrees in neuroscience and dance from the University of Florida. Beyond Live Science, Lanese's work has appeared in The Scientist, Science News, the Mercury News, Mongabay and Stanford Medicine Magazine, among other outlets. Based in NYC, she also remains involved in dance and performs in local choreographers' work.

 Live Science Plus
Live Science Plus




