Inherited diseases don't work like we thought they did
"Monogenic" diseases, triggered by mutations in just one gene, may actually be more complex than scientists thought.
Get the world’s most fascinating discoveries delivered straight to your inbox.
You are now subscribed
Your newsletter sign-up was successful
Want to add more newsletters?
Join the club
Get full access to premium articles, exclusive features and a growing list of member rewards.
For decades, scientists have delved into the genetic causes of disease by studying patients with those diseases and their families, picking through their genomes in search of genetic mutations that could be the cause.
It's a method that has turned up hundreds of mutations, many thought to be responsible for diseases in almost 100% of the people carrying them. Such mutations have been linked to myriad conditions, from thyroid cancer to ovarian insufficiency to certain forms of diabetes.
But new research is finding that these so-called "monogenic diseases" actually aren't caused by single gene mutations at all. In fact, plenty of perfectly healthy people are walking around with these mutations and no sign of disease, population-based studies are revealing.
"It kind of challenges our standard dogma," said Caroline Wright, a professor of genomic medicine at the University of Exeter in England. Wright has found gene variants that seem to cause disease all the time in patient samples but only in a minority of people in the general population.
"In much of single-gene genetics we've often assumed that a particular genetic cause is necessary and sufficient, and everything else is irrelevant," Wright told Live Science. "And what we're seeing is that that's not necessarily true."
It's a finding that impacts both genetic counseling for patients with family histories of genetic disease and possible treatments for these disorders.
A complicated inheritance
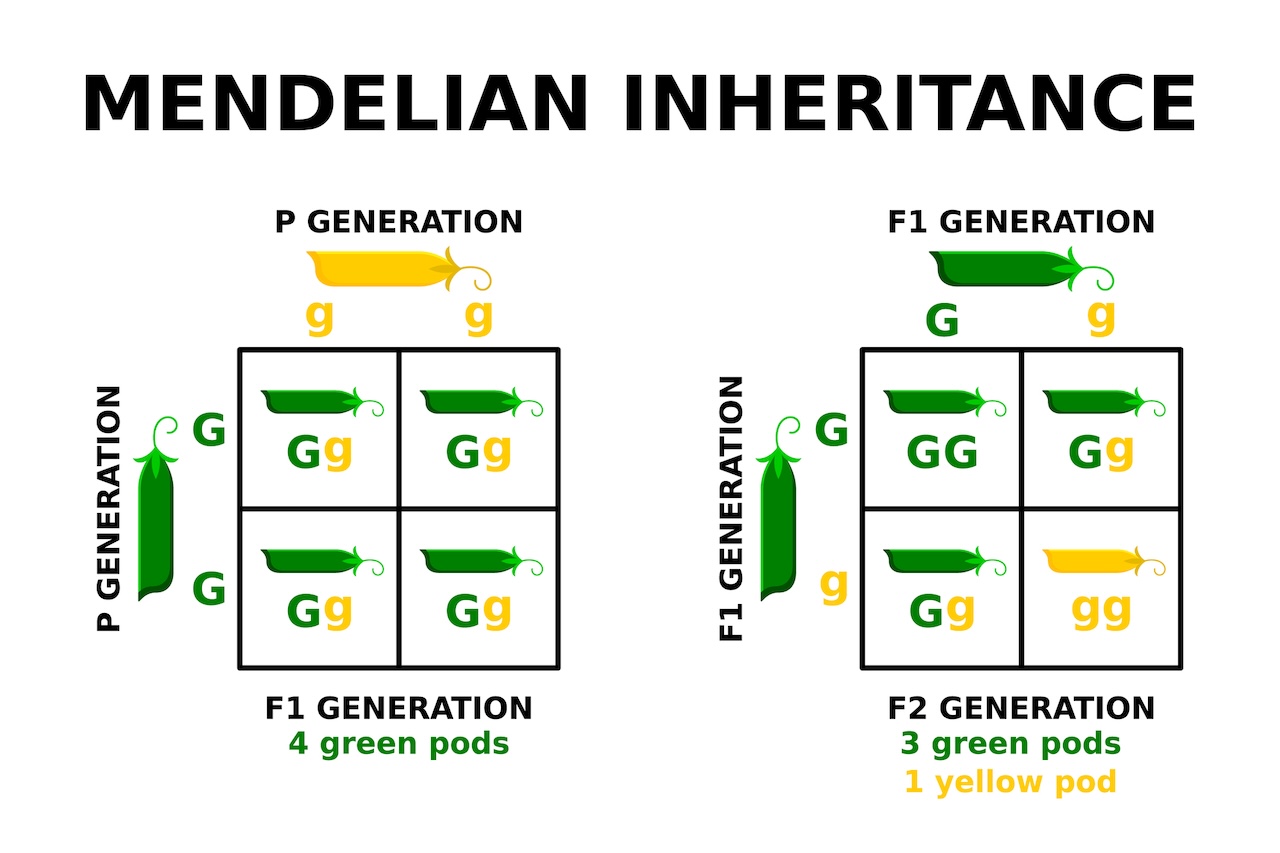
In the mid-1800s, Gregor Mendel worked out the rules of inheritance with the help of pea plants, establishing the basics of modern genetics: Offspring receive a copy of each gene from each parent. In some cases, a gene is dominant, meaning just one copy is enough to ensure that the gene's instructions are expressed in the body. In others, it is recessive, meaning two copies are needed.
Get the world’s most fascinating discoveries delivered straight to your inbox.
Beyond those basics, things get more complex, as not all genes are as simple as the "brown eyes dominant, blue eyes recessive" many people learn in middle school science class. Genes interact with one another and with environmental factors, and those interactions determine the person's phenotype, or traits. In practice, the likelihood that a person with a particular genotype, or combination of genes, expresses a particular phenotype is known as "penetrance."
Some diseases, known as monogenic diseases, have long been classified as having 100% penetrance, meaning people with a given gene mutation always get the disease. The lethal nervous system disorder Tay-Sachs disease, for example, is a recessive condition that develops in all babies with two copies of a particular mutated gene, one inherited from each parent.
Other conditions, such as Crohn's disease and schizophrenia, are classified as polygenic, meaning they arise from the interactions of hundreds to thousands of genes, as well as environmental factors. In polygenic conditions, there isn't a single genetic switch that determines if the person has the disease or not — rather, scientists calculate risk scores that try to take into account as much of the person's genetic spectrum as is understood. The higher the score, the higher the probability of disease.
Many rare diseases were once thought to be monogenic, on the same spectrum as Tay-Sachs. But now, research is showing they are more akin to schizophrenia. What's cracked this new knowledge open is the creation of huge genetic databases from healthy populations.
When gene sequencing was expensive, researchers were limited to looking at the genomes of people with diagnosed diseases and their families. Take inherited retinal degenerations, a constellation of conditions in which the layer of the eye containing light-sensing cells degrades, leading to early vision loss. By testing patients and their families, researchers found genes that were more common in people with the disorder and genes passed down in the family line that raised the risk of disease. Not all of these genes conferred a 100% risk of disease in these clinical samples, but many appeared to come close.
But that method suffers from what's known as "ascertainment bias," said Dr. Eric Pierce, director of the Ocular Genomics Institute at Mass Eye and Ear and an ophthalmologist at Harvard Medical School. Because you're only looking at people with the disorder, you may indeed see gene variants that are common in people with the disorder. But what you can't see is whether healthy people also carry the same variants.
Today, gene sequencing is cheap, and huge studies like the U.S. National Institutes of Health All of Us cohort and the U.K. Biobank collect genetic data and medical records from hundreds of thousands or millions of individuals, sometimes following them through time. These individuals are part of the "general population," meaning they are not specifically included in the cohorts because they have a particular medical condition.
Pierce and his colleague Dr. Elizabeth Rossin, an ophthalmologist and vitreoretinal surgeon at Mass General Hospital and Harvard University, looked in these databases for 167 gene variants thought to lead to severe vision loss almost all the time. What they found was that, in actuality, people with these gene variants had vision loss less than 30% of the time.
"That means for that other 70% of people there is something about the rest of their genome or environment that is changing the way they manifest," Rossin told Live Science.
The supporting cast
If the genes linked to the condition are lead actors, the rest of the genome and the environment are the supporting cast.
In patients and their relatives, the leading-actor genes tend to be shared. But so does the supporting cast. That makes the role of those secondary genes difficult to tease out. In the broader population, the supporting casts aren't the same, so scientists can start to probe the role of the rest of the genome in either protecting against disease or making a disorder more likely to emerge.
There is now a long list of single-gene variants that seem to be necessary to cause a disease, but not sufficient: They appear in almost everyone with the disease, but at the same time, don't seem to trouble most the people that carry them. For example, Wright and her colleagues have found that genetic variants that seemed to cause thyroid cancer in 95% of clinical populations only lead to disease in between 2% and 19% of the general population.
One study by Wright, not yet peer-reviewed, found that variants thought to cause "brittle bone disease," or osteogenesis imperfecta, almost 100% of the time may only cause the disorder about 21% to 40% of the time. Another finds similar results with variants thought to cause a rare childhood eye cancer. Other researchers have turned up the same patterns in mitochondrial diseases, certain inherited forms of diabetes, and ovarian insufficiency, a condition that causes early menopause.
On the flip side, sometimes these studies turn up more risk than expected in the general population. Huntington's disease is a degenerative neurological condition caused by a repeating genetic sequence in the Huntington's gene. It was initially thought to be inherited in an autosomal dominant pattern, meaning if you inherited at least one copy of the defective gene from your parents, you always got the disease.
Just because you can demonstrate that your variant affects a process [in the lab] doesn't actually necessarily mean that that's what is happening in your particular cell in that system.
Anna Murray, professor of genetics at the University of Exeter
Later research showed that the disease was somewhat dose-dependent; when people have 40 or more of these genetic repeats, they eventually develop the disease. But a 2016 study found that 1 in 400 people in the population carry 36 to 39 repeats of the gene — right on the edge of the threshold. Not all these people will develop Huntington's, but they are at higher risk, said Michael Hayden, a professor of medical genetics at the University of British Columbia who led that research.
This discovery led Hayden and his team to further study patients with Huntington's who carry 36 to 39 genetic repeats. They found that those at the highest risk of developing the disease earlier in life carry an additional variant nearby that essentially makes the gene act as if the repeated segment were longer.
The Huntington's research is an example of how population and patient samples can complement each other to lead scientists to new answers about how diseases develop.
In ovarian insufficiency, where more than 99% of genetic variants thought to cause the disease were also present in asymptomatic women, population studies also point to the need for better basic cellular research on the condition, said Anna Murray, a professor of human genetics at the University of Exeter in England who led that work. Many genes involved in that condition have multiple roles in the body and multiple interactions with other genes in tissues beyond the ovary, she said.
"Just because you can demonstrate that your variant affects a process [in the lab] doesn't actually necessarily mean that that's what is happening in your particular cell in that system," Murray told Live Science.
Individual risk
Studies of patient groups can reveal an upper bound on the risks of a genetic variant, Wright said, while population-level studies can provide a lower bound. The challenge is helping patients understand what this range means for their own personal risk — a question that researchers are trying to answer.
As genetic screening becomes more ubiquitous, understanding the meaning of individual variants is increasingly important. Parents doing IVF and screening embryos for health conditions, for example, might make very different decisions about which embryos to implant if they are told the risk of genetic disease from a particular variant is 100% versus 20%. Likewise, people getting genetic counseling need to understand if their risk of disease is truly as high as studies on patient populations would suggest, or if they have protective factors that bring their risk down.
The findings may also help refine gene therapy treatments that target specific disease-causing genes. Because these genes do cause the disease in some people, these gene therapies will continue to be important, Pierce said. But understanding the rest of the genetic milieu could help boost the effectiveness of such therapies.
"We might be able to predict more accurately who will respond best to genetic therapies," Pierce said. "We might also identify novel targets for therapies following the identification of the additional genetic factors that influence disease expression."
Rossin and Pierce are now working on large global collaborations to learn more about what modifies the genetic risk in retinal disorders. Other researchers are looking to do the same for disorders such as ovarian insufficiency.
Right now, there are limited treatment options for many of the conditions under study. That means that there may be few options for prevention or early treatment, even if the genes fueling the conditions are better understood.
But as more new treatment options emerge, it will be increasingly important to understand the nuances of risk for individual patients, Hayden said. Understanding that risk could help patients make decisions about preventative treatment.
"When therapies are available for those diseases, early treatment—and particularly early treatment for degenerative disease of the brain and the eye—is better than later, because you can't replace neurons," Hayden said. "So you'll want to know your risk."

Stephanie Pappas is a contributing writer for Live Science, covering topics ranging from geoscience to archaeology to the human brain and behavior. She was previously a senior writer for Live Science but is now a freelancer based in Denver, Colorado, and regularly contributes to Scientific American and The Monitor, the monthly magazine of the American Psychological Association. Stephanie received a bachelor's degree in psychology from the University of South Carolina and a graduate certificate in science communication from the University of California, Santa Cruz.
You must confirm your public display name before commenting
Please logout and then login again, you will then be prompted to enter your display name.

 Live Science Plus
Live Science Plus