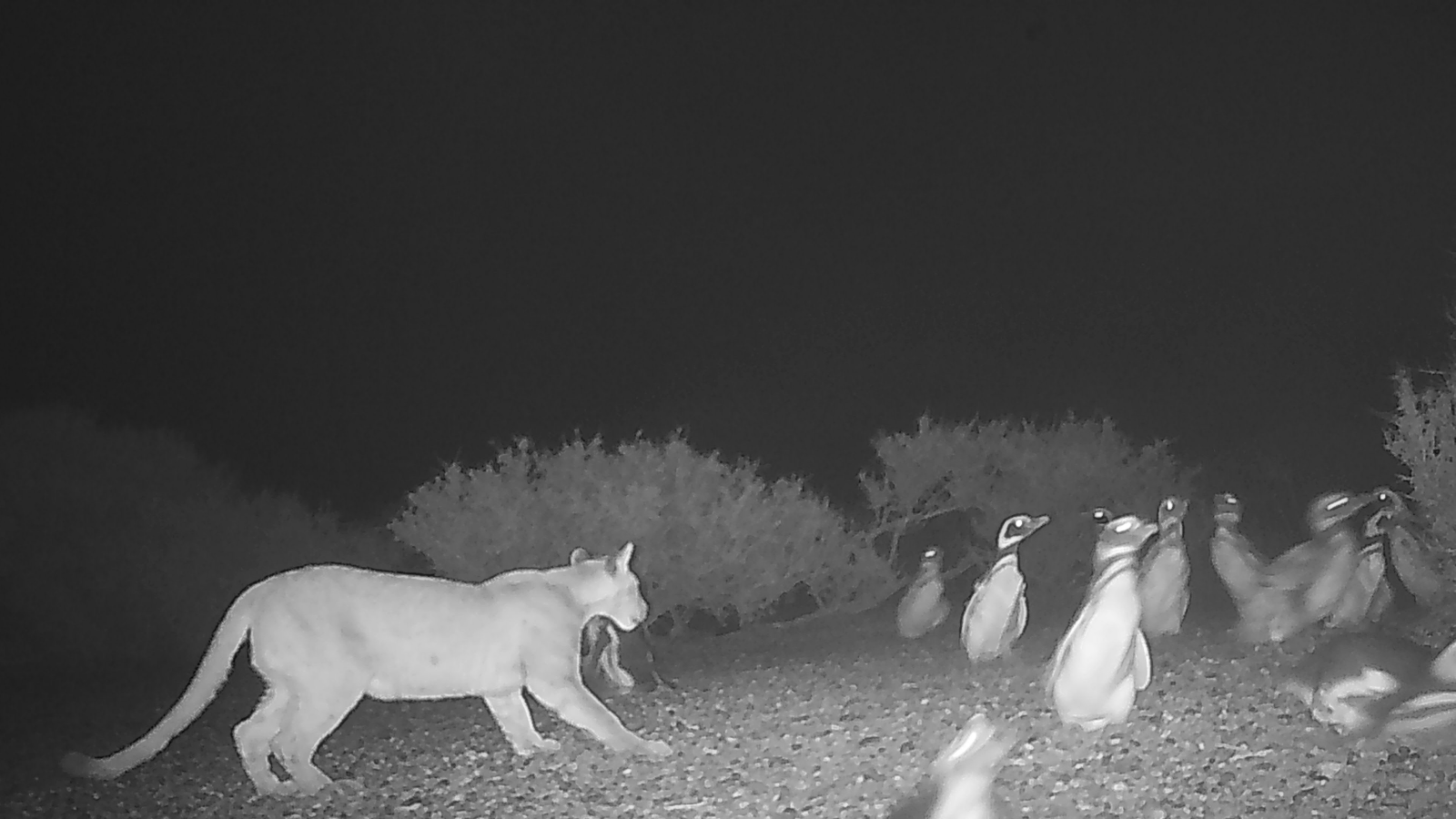
How a Cell Knows Friend From Foe

This article was provided by the National Institute of General Medical Sciences (NIGMS), part of the National Institutes of Health. NIGMS supports basic research that increases understanding of biological processes and lays the foundation for advances in disease diagnosis, treatment and prevention. Carolyn Beans is a science writer for NIGMS. This article was provided to Live Science's Expert Voices: Op-Ed & Insights.
Cells are faced with many decisions: When's the best time to produce a new protein? To grow and split into two? To treat another cell as an invader? Scientists are working to understand how cells make these and many other decisions, and how these decisions contribute to health and disease.
The ability of an organism to distinguish its own cells from those of another is called allorecognition, and it is an active area of research. Immune cells use a system called the major histocompatibility complex (MHC) to identify which cells belong to the body and which are foreign. Brain cells, skin cells and nearly all other cells in our bodies have MHC proteins on their outer surfaces. Immune cells use those protein markers to decide whether other cells belong, or if they should be attacked.
But the system isn't perfect. An invading pathogen might go undetected — the hepatitis C virus can evade immune cells for years. Or, the body might mistake its own cells for intruders, leading to autoimmune diseases like lupus and inflammatory bowel disease.
An early step in developing more targeted approaches to address these issues is to gain a better understanding of the molecular mechanisms involved in allorecognition. "At a basic level, we are still trying to understand how one cell recognizes another," says Gad Shaulsky of Baylor College of Medicine.
Shaulsky is one of many researchers working to figure this out. Because allorecognition in human cells involves a dizzying number of protein interactions, Shaulsky and his team study a simpler creature, the soil amoeba Dictyostelium discoideum.
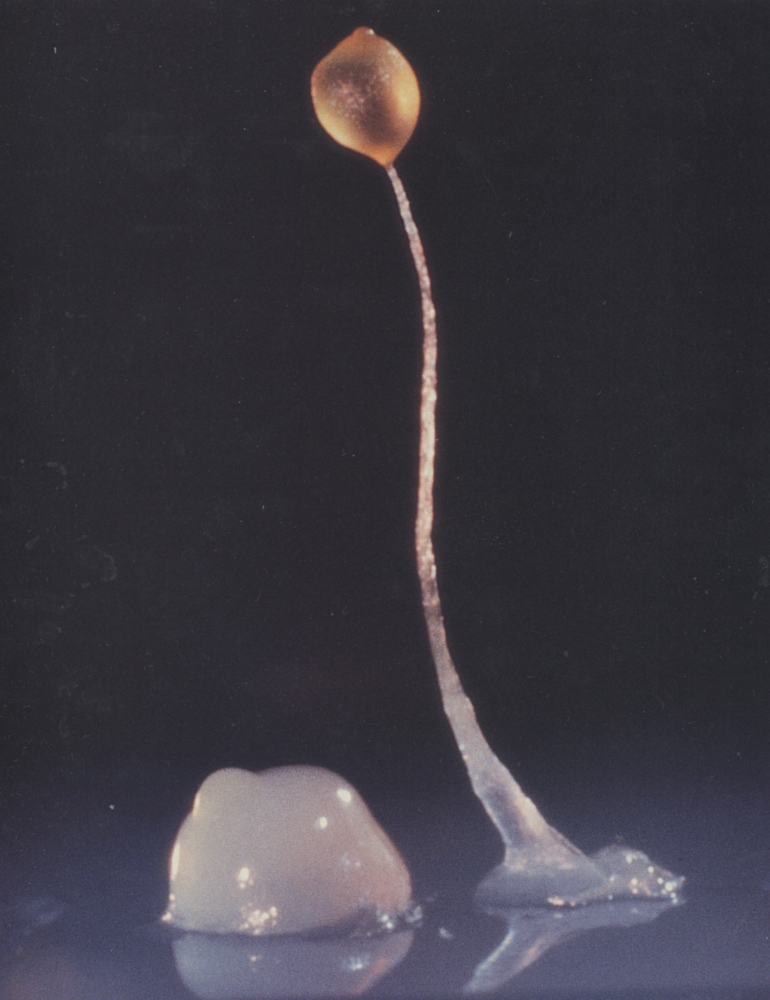
"When amoeba cells have abundant food, they behave as unicellular organisms that don't cooperate with one another," Shaulsky explains. "When you starve them, they aggregate with their close relatives into a fascinating structure of about 100,000 cells that has coordinated movement and can sense light and heat even though it doesn't have a nervous system."
Get the world’s most fascinating discoveries delivered straight to your inbox.
Using a series of experiments that entailed inserting, deleting and swapping amoeba genes, Shaulsky determined that amoebas use two sets of proteins, TgrB1 and TgrC1, to recognize cells from the same strain. An amoeba cell has a copy of each protein protruding from its outside membrane.
Different strains of amoebas have different versions of these proteins, so when two amoeba cells from the same strain meet, the TgrB1 proteins from each cell lock into the TgrC1 proteins in the other cell, enabling the cells to join together. When cells from different strains meet, their proteins don't match, so they can't aggregate.
By conducting additional gene-swapping experiments, Shaulsky now wants to learn exactly what happens inside an amoeba cell, on a molecular level, after the two proteins connect. He thinks that contact between the proteins might set off a cascade of signals that ultimately tells cells whether or not to join together with a close relative.
The Tgr protein system in the amoeba is similar to our own MHC system, but Shaulsky is quick to point out that these allorecognition processes evolved independently. The different origins mean that the molecular mechanisms he uncovers in the amoeba will not necessarily be the same in humans.
However, gaining new insights into how allorecognition works in this simple creature may inform allorecognition research in more complex organisms, including humans.
Follow all of the Expert Voices issues and debates — and become part of the discussion — on Facebook, Twitter and Google+. The views expressed are those of the author and do not necessarily reflect the views of the publisher. This version of the article was originally published on Live Science.



