How does DNA know which job to do in each cell?
If each cell carries the same blueprint, what sets them apart?


A copy of your DNA is harbored in the nuclei of all 37.2 trillion of your cells. Theoretically, all of these cells have the same capabilities, because they carry the same blueprint. So how does your DNA know when it's in a blood cell versus an olfactory cell, for example? How does it know which genes need to be "switched on"? How does a cell know and carry out its function?
Like all things DNA-related, it is a multifactorial and highly regulated process. In humans and other organisms with eukaryotic cells (which have an enclosed nucleus), a concept known as "central dogma" explains how DNA serves as an instruction manual, with DNA informing messenger RNA (mRNA), which is then used as a road map for protein production. So, transcribing the right piece of DNA into mRNA is just the first step in ensuring the cell has all the proteins it needs.
A special protein called a transcription factor turns on genes, said Karen Reddy, an assistant professor of biological chemistry at John Hopkins University School of Medicine. The transcription factors bind to the DNA to increase or decrease the expression of specific genes. But it raises a question: Where does the transcription factor come from?
Related: Are you genetically more similar to your mom or your dad?
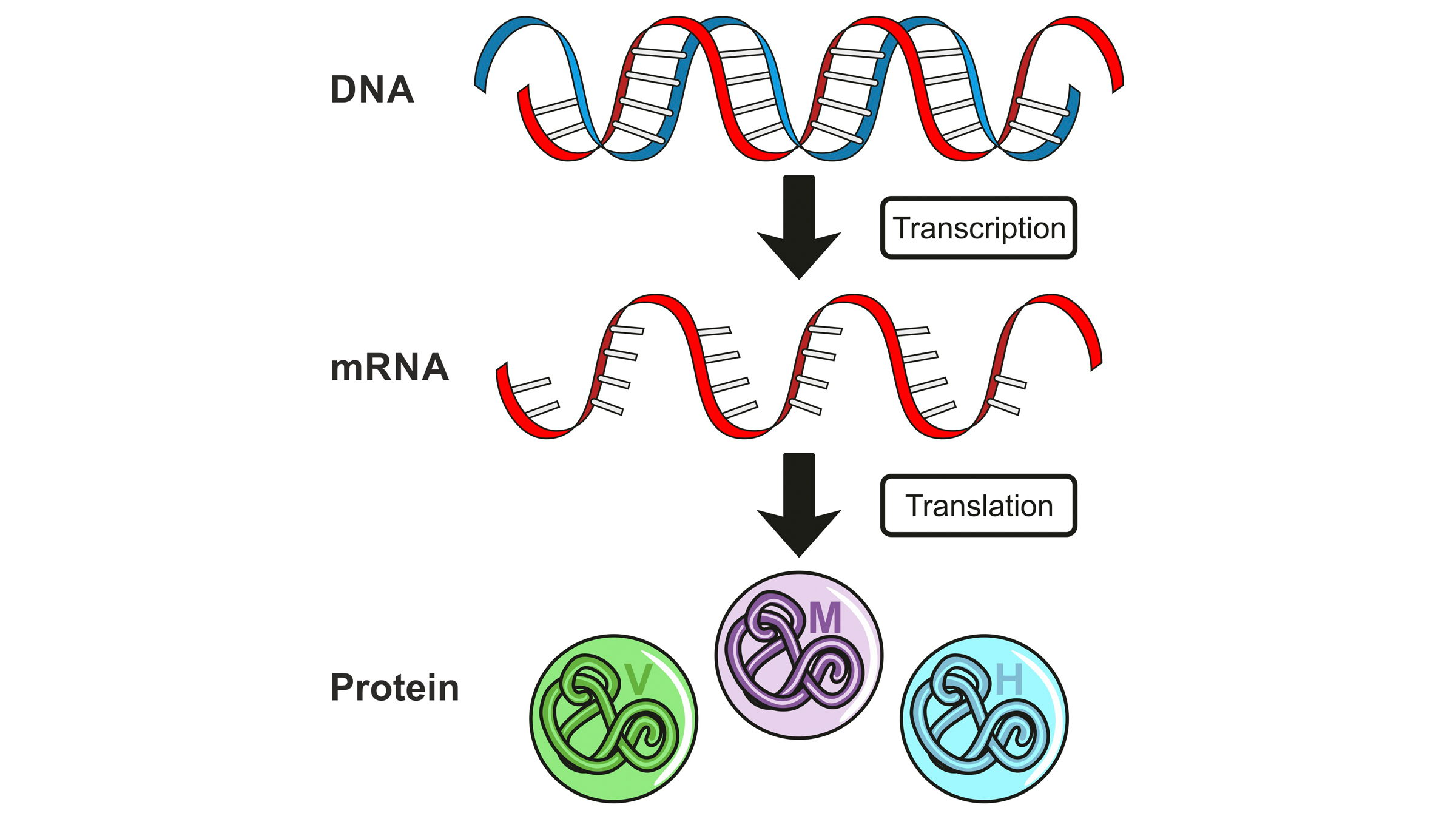
This illustration shows "central dogma," with DNA informing messenger RNA (mRNA), which serves as a roadmap for protein production in cells.
"Lots of transcription factors are reused from cell to cell to cell," Reddy told Live Science. It's like how the same parts can be used in different cars. One transcription factor can activate different genes in different cell types. For instance, a transcription factor used in olfactory cells called Olf-1 is identical to the one used in specifying B cells, Ebf-1. And the transcription factor knows to activate different genes in these cells because the DNA is organized and packaged differently in different cell types, also known as having different chromatin landscapes.
In the nucleus, a complex of DNA, proteins and RNA function together to package the long strands of DNA. The complex is called chromatin. How the DNA is wrapped around a complex of proteins called histones, and the chemical modifications to those histones, is called the chromatin landscape. This impacts which genes are more or less exposed. In a given cell type, some genes are poised for activation by the transcription factor because of how they are exposed in the chromatin structure, Reddy said. Others are repressed — or tucked away — by the chromatin landscape. These can still be turned on, but first there need to be enough transcription factors and chromatin modifiers to alter the chromatin structure and reveal them.
"There's cross talk between the chromatin landscape and the transcription factor universe," Reddy said.
Get the world’s most fascinating discoveries delivered straight to your inbox.
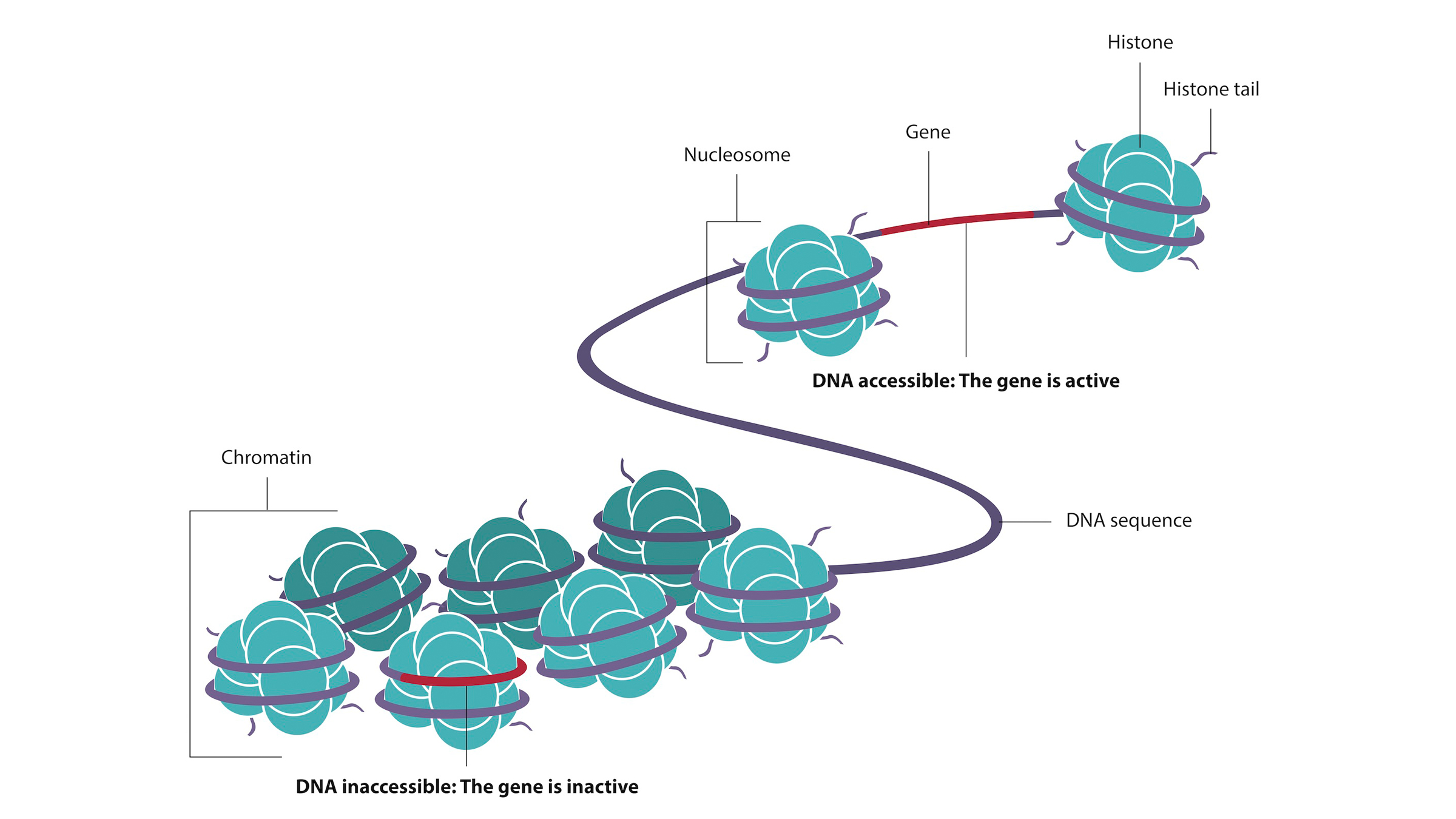
DNA wraps around proteins called histones to form chromatin.
Overlaying both of these factors is the 3D architecture in the cell nucleus, Reddy said, or how the chromatin is folded and organized in the nucleus. This folding facilitates interactions between the genes that need to be expressed and elements that increase their expression. The active — or needed — parts of DNA in a given cell type are grouped near the center, while the inactive sections are close to the outside of the nucleus.
Some elements that control how a gene is expressed, like a promoter that can turn the gene on or off, are immediately nearby the gene. But other elements, like a tissue-specific enhancer that increases gene expression, can be much farther away from the gene they need to enhance for the cell. The folding or 3D architecture brings the enhancer in proximity to the gene of interest, Reddy said.
Finally, there are processes that make more long-lasting changes to the DNA itself. For example, DNA methylation involves adding a methyl group to a nucleotide (the DNA "building block" cytosine and its backbone) and is generally associated with repression of a gene, Reddy said. DNA methylation can be transmitted from generation to generation, influences which genes are turned on or off in a specific type of cell, and keeps you from over-expressing certain genes, which, in turn, can lead to conditions such as nervous disorders and cardiovascular disease, according to a 2015 review in the journal Cureus.
RELATED MYSTERIES
All of these levels — DNA methylation, chromatin landscape, folding and transcription factors — are important regulatory steps for expressing essential genes in the right place at the right time, Reddy said. "Any of these levels of control are perturbed in a disease like cancer."
The good news is that these regulatory elements back each other up. "You can have something go kind of wrong, and you'll be OK as a cell, because these processes reinforce one another," Reddy said.
Originally published on Live Science.

Donavyn Coffey is a Kentucky-based health and environment journalist reporting on healthcare, food systems and anything you can CRISPR. Her work has appeared in Scientific American, Wired UK, Popular Science and Youth Today, among others. Donavyn was a Fulbright Fellow to Denmark where she studied molecular nutrition and food policy. She holds a bachelor's degree in biotechnology from the University of Kentucky and master's degrees in food technology from Aarhus University and journalism from New York University.

 Live Science Plus
Live Science Plus





