Going Viral: When Living Networks Go Awry
Networks — both real and virtual — are everywhere, from our social media circles to the power grid that delivers electricity. The interactions of genes, proteins and other molecules in a cell are examples of networks, too. Scientists working in a field called systems biology study and chart these and other living networks to learn how the individual parts work together to make a functioning whole and what happens when these complex, dynamic systems go awry.
Just as you can't understand how a social media post "goes viral" by observing the activity of individual users, scientists can't fully understand the behavior of a cell or other biological system by studying its individual components in isolation. But, by combining data from experiments in living systems with powerful computer modeling techniques, scientists can explore questions that might be difficult or impossible to answer through traditional lab experiments alone — such as what drives a cancer cell to grow into a life-threatening tumor.
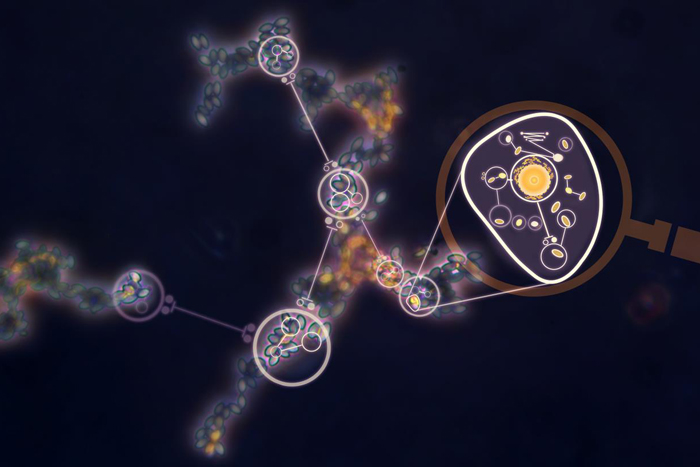
According to Aaron Brooks, a biologist at the National Institutes of Health-funded Institute for Systems Biology in Seattle, networks typically are illustrated as diagrams consisting of "nodes" and "edges." The nodes, often shown as circles, represent the objects in the network. The edges, often shown as lines, depict the interactions between those objects.
For example, the network diagram in the image above depicts yeast cells (superimposed circles) and the biochemical "chatter" between them (lines) that tells the cells to gather together in clumps. This clumping helps them survive stressful conditions like a shortage of nutrients.
Network diagrams provide more than just hub-and-spoke pictures, Brooks explains. They can yield information that helps us better understand — and potentially influence — complex phenomena that affect our health. Drug interaction networks can predict and help prevent harmful combinations of medications. Gene interaction networks can shed light on the basis for complex traits like disease susceptibility. And social interaction networks can help predict how an infectious disease may spread and point to ways to contain it.
"Network dynamics fascinate us," says Chris Lausted, a senior research engineer at ISB. "To survive, biological systems need to be very sensitive to some subtle environmental signals while tolerating huge changes in others. Mapping these networks helps us understand which changes cause disease and what can be done to restore health."
At the USA Science & Engineering Festival in Washington, D.C., last month, Brooks and another ISB biologist, Martin Shelton, introduced thousands of students to some of these concepts. Through a hands-on exhibit sponsored by NIH, the students first tinkered with a small, four-node network wired with optical cables and lights. The initial challenges were simple: establishing long-distance communication with a friend or understanding how germs and the immune system interact to affect a person's health. By removing individual nodes or links in their networks, participants could see how the changes affected the network overall.
Get the world’s most fascinating discoveries delivered straight to your inbox.
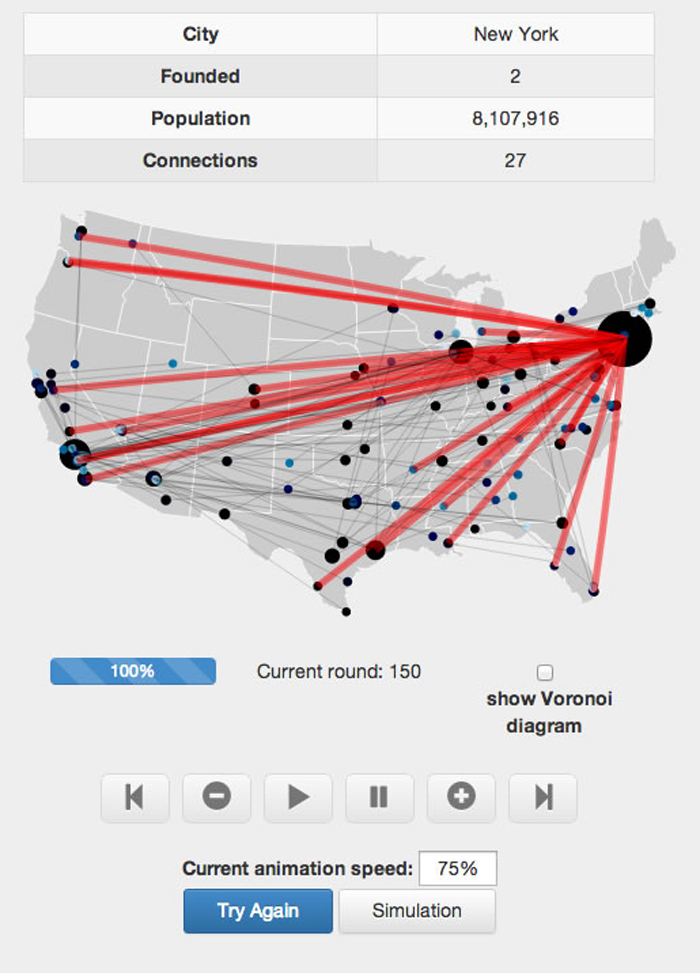
The next challenge was much bigger: designing a communication network for 150 U.S. cities. The systems were scored according to how well they could withstand a random failure in one of the cities (a trait systems scientists call "robustness"). After observing the structure and performance of their systems, the junior network engineers could see how their design choices compared to those of other participants and repeat the activity to earn a higher score. The combination of activities gave students a feel for the real-world approaches, tools and reasons for studying complex systems.
"There was a point, while walking the kids through the activities, where their eyes would light up and they would smile, and you knew that they understood," said Shelton. "I think many scientists became scientists because they fell in love with that very same feeling as a kid — the joy of learning something new that changes your perspective and understanding of the world around you."
The research reported in this article was funded in part under NIH grant P50 GM076547.
Elia Ben-Ari also contributed to this story.
This Inside Life Science article was provided to Live Science in cooperation with the National Institute of General Medical Sciences, part of the National Institutes of Health.
Learn more:
National Centers for Systems Biology
Also in this series:
Cells Are Chatty: Here's What They May Be Saying

 Live Science Plus
Live Science Plus






