Researchers to Harness the Power of Molecular Self-Assembly
This Behind the Scenes article was provided to LiveScience in partnership with the National Science Foundation.
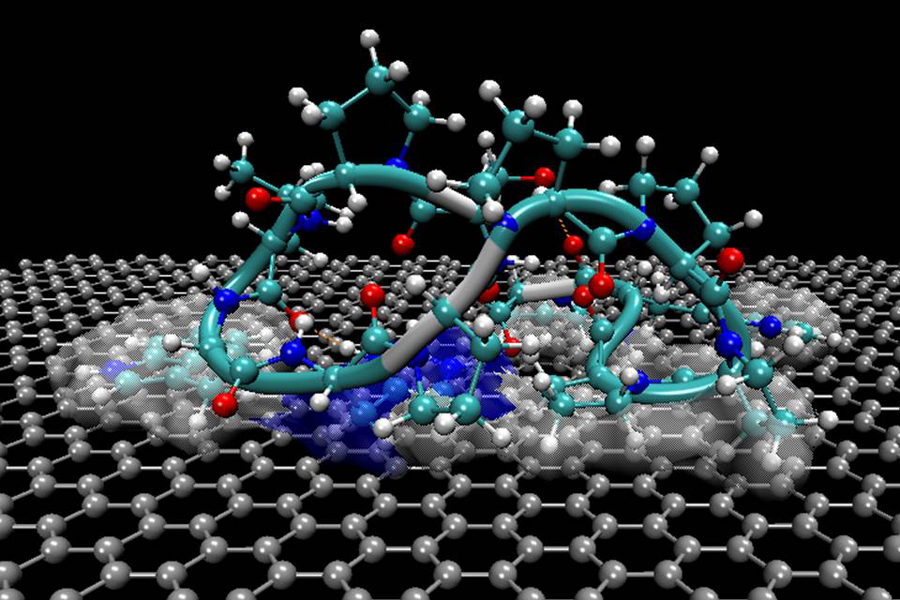
Researchers at the University of Washington have demonstrated how peptides, or short chains of amino acids, assemble by themselves into nano-sized structures on solid surfaces such as graphite and other layered minerals.
These findings are expected to help researchers harness the power of molecular self-assembly — the process by which molecules form a defined, well-organized arrangement without interference from external sources.
Molecular self-assembly " . . . gives a tremendous power to the scientist to make controlled nanostructures — the hallmark of nanotechnology," said Mehmet Sarikaya, professor of materials science and engineering at the university and director of the NSF-funded Genetically Engineered Materials Science & Engineering Center.
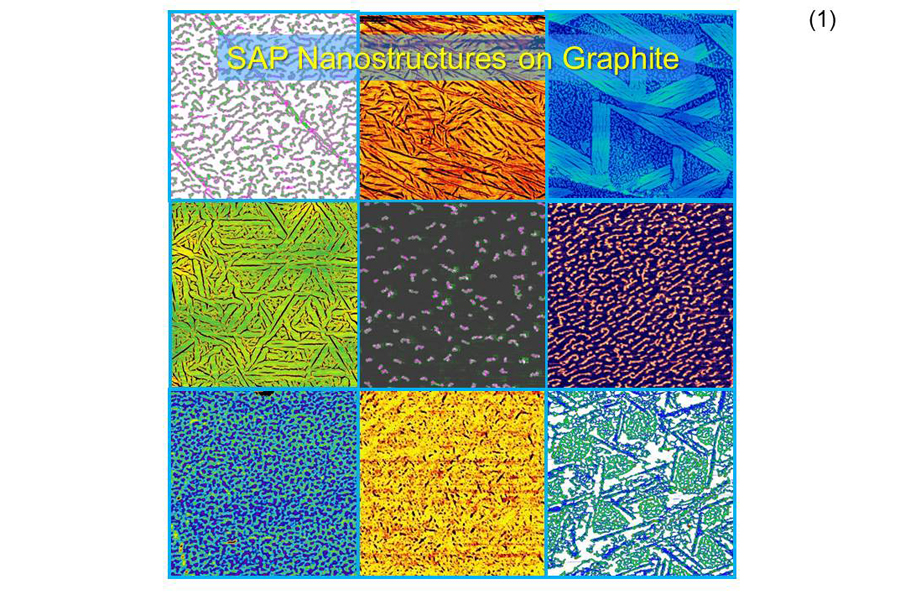
Controlling Self-Assembly
Sarikaya's research involved observing selected amino acids arrange themselves into a linear form, and then fold and coil into a 3-D protein. These observations were conducted at regular intervals ranging from 10 seconds to 15 hours in order to capture the progression of events.
This research was conducted with atomic force microscopy. AFM involves using high-resolution microscopes to produce images down to the molecular and atomic levels.
Get the world’s most fascinating discoveries delivered straight to your inbox.
Analyses of the Sarikaya's observations revealed which amino acids apparently control surface and inter-molecular interactions of peptides that led to their self-assembly. Based on these insights, Sarikaya was able to control the self-assembly and formation of specific biomolecular nanostructures on graphite surfaces; these nanostructures were dubbed as self-assembled peptides.
The results of Sarikaya's research will advance efforts to use molecular self-assembly to engineer nanoscale machinery and microelectronics that are incorporated into:
- Biomolecular nanosensors, which may be used in molecular probing for cancer targets.
- Nanophotonics devices, such as self-assembled Light-Emitting Diodes, which are light sources used in many applications ranging from general lighting to aviation lighting
- Biofuel cells, which mimic bacterial interactions in nature that produce electrical currents
- Bioelectronics, which use electrical stimuli to manipulate various biological systems
Additional research of protein self-assembly and protein interactions that are related to this research may also aid in drug design. "Big Pharma companies cannot easily design drugs because many of these interactions and the resultant structures are not known," said Sarikaya. "Short peptides assembling on solid surfaces … may be a way to overcome some of the design and assembly problems encountered ..."
Editor's Note:The researchers depicted in Behind the Scenes articles have been supported by theNational Science Foundation, the federal agency charged with funding basic research and education across all fields of science and engineering. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author and do not necessarily reflect the views of the National Science Foundation. See the Behind the Scenes Archive.

 Live Science Plus
Live Science Plus






