DeepMind says it can predict the shape of every protein in the human body
And in 20 other animals often studied by science, too.

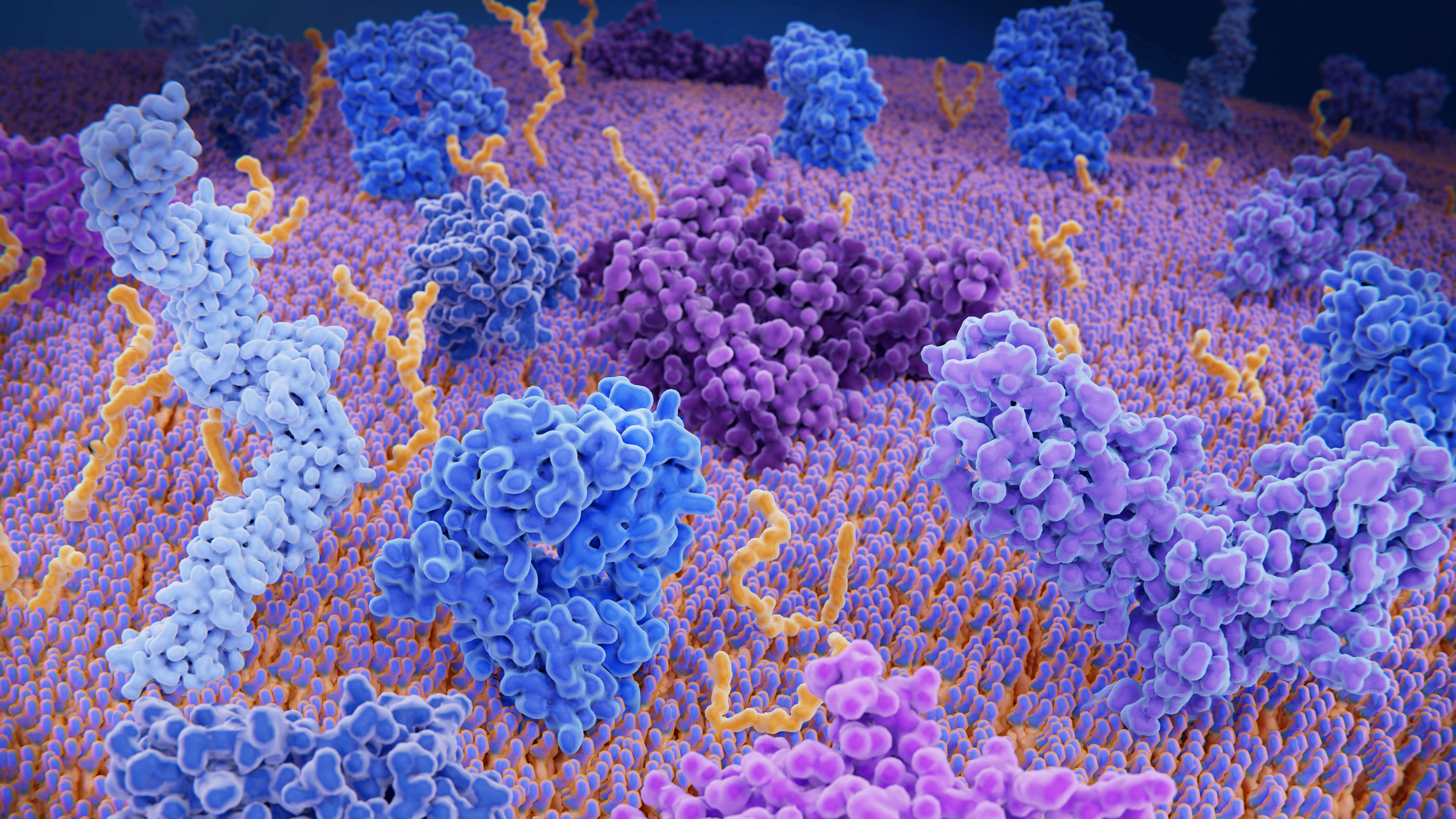
Proteins are complex structures in the body. Here, the brightly colored and twisty blobs represent different immune system proteins on the outer layer of a T-cell, a type of white blood cell that helps the body to identify foreign invaders.
The artificial intelligence (A.I.) company DeepMind says it will soon release a database of the shape of every protein known to science — more than 100 million.
That's every structured protein in the human body, as well as in 20 research species, including yeast and E. coli bacteria, fruit flies and mice. Prior to the company's AlphaFold project, which uses artificial intelligence to predict protein shapes, only 17% of the proteins in the human body had their structures identified, according to Technology Review.
"It looks astonishingly impressive," Tom Ellis, a synthetic biologist at Imperial College London, told Technology Review.
Protein folding is incredibly complex. Proteins are made of long strands of building blocks called amino acids, which wrap themselves into strange and complicated shapes to form functional structures. Unraveling these structures in the laboratory takes a long time, but DeepMind announced in December that its AlphaFold algorithm can determine the shape of proteins down to the atom in minutes. So far, AlphaFold has predicted 36% of human proteins with atomic-level accuracy, and has predicted more than half with accuracy good enough to spark research on the proteins' functions, according to the company. (About a third of the proteins in the body don't have a structure unless they bind to something else, so DeepMind can't accurately predict their shapes.) AlphaFold makes these predictions using a neural network, a type of algorithm meant to mimic how the brain processes information, and which is particularly good at recognizing patterns — such as how particular sequences of amino acids interact — in large amounts of data.
The predicted shapes still need to be confirmed in the lab, Ellis told Technology Review. If the results hold up, they will rapidly push forward the study of the proteome, or the proteins in a given organism. DeepMind researchers published their open-source code and laid out the method in two peer-reviewed papers published in Nature last week.
RELATED CONTENT
They have now made about 350,000 protein structures freely available in the AlphaFold Protein Structure Database, according to a company announcement. These include the 20,000 or so proteins expressed by the human genome. (When proteins are "expressed," that means that information stored in the genome gets converted into instructions to make proteins, which then perform some function in the body.) In the coming months, the company plans to add almost every sequenced protein known to science.
Understanding protein structure can help researchers delve into the causes of diseases and enable them to discover new drugs that will carry out a particular function in the body. According to DeepMind, researchers are already using AlphaFold's discoveries to study antibiotic resistance, to study the biology of the SARS-CoV-2 virus, which causes COVID-19, and to seek new enzymes that can be used to recycle plastics.
Get the world’s most fascinating discoveries delivered straight to your inbox.
Originally published on Live Science

Stephanie Pappas is a contributing writer for Live Science, covering topics ranging from geoscience to archaeology to the human brain and behavior. She was previously a senior writer for Live Science but is now a freelancer based in Denver, Colorado, and regularly contributes to Scientific American and The Monitor, the monthly magazine of the American Psychological Association. Stephanie received a bachelor's degree in psychology from the University of South Carolina and a graduate certificate in science communication from the University of California, Santa Cruz.