5 Ways Computers Boost Drug Discovery

Get the world’s most fascinating discoveries delivered straight to your inbox.
You are now subscribed
Your newsletter sign-up was successful
Want to add more newsletters?
Join the club
Get full access to premium articles, exclusive features and a growing list of member rewards.
High-Tech Drug Design

The design of airplanes, bridges and even heart stents starts with computer-generated models that not only detail what the products could look like, but how they'd work under different conditions. In recent years, scientists have started to use the same approach to help design new drugs.
Here are a few examples of how computation is contributing to many aspects of the drug discovery process, including identifying promising compounds for further testing in the lab and in clinical trials.
Learn more:
This Inside Life Science article was provided to LiveScience in cooperation with the National Institute of General Medical Sciences, part of the National Institutes of Health.
More Complete Models
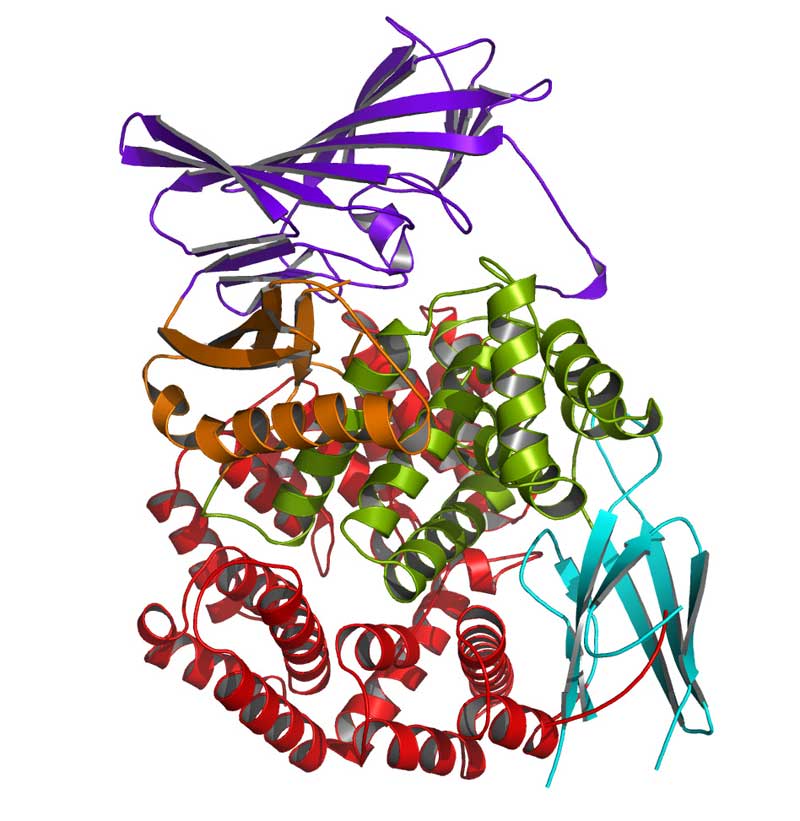
Most drugs work by either blocking or stimulating the activity of specific proteins in the body. Pain relievers, for example, block an enzyme involved in inflammation. To create a drug that will interact with a protein target in the desired way, chemists typically start with a computerized structural model of the protein bound to a natural molecule that "unlocks" a biological action. Then, they try to design small molecules that behave like the natural one. But this approach is only as good (and as accurate) as the starting protein model.
Researchers at the University of Texas at Austin recently improved algorithms for modeling short sections of a protein’s structure. By capturing additional details of a protein's shape, they can better understand, identify and predict how a potential drug molecule would bind.
Forecasting New Uses
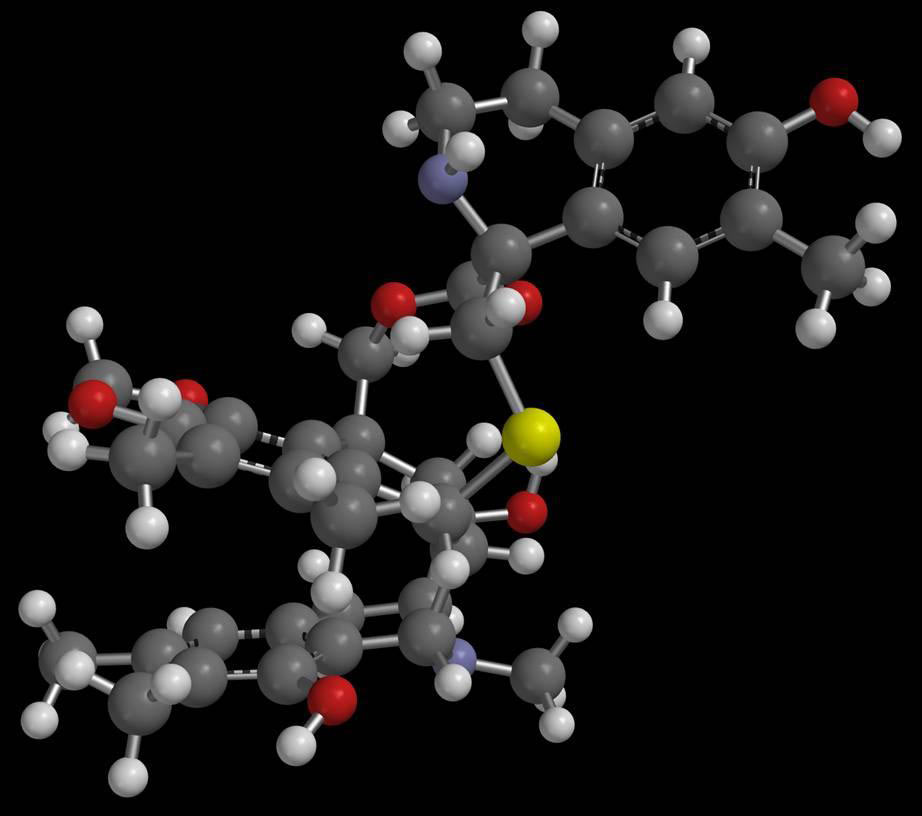
Developing a new drug and bringing it to market can take 15 years and cost more than $1 billion. An alternative is to identify and test FDA-approved drugs for new uses, also called drug repositioning. By computationally sifting through public databases of genomic information, Stanford University researchers have matched 53 human diseases, including cancers, Crohn's disease and cardiovascular conditions, to existing drugs that might work as treatments for them.
Get the world’s most fascinating discoveries delivered straight to your inbox.
The scientists confirmed some already known matches — validating the usefulness of the approach — but they also identified some surprising pairs. For instance, topiramate, an anticonvulsant used to treat epilepsy, emerged as a good match for inflammatory bowel disease. The finding held up when tested on rat and mouse models.
Drug Screening Tools
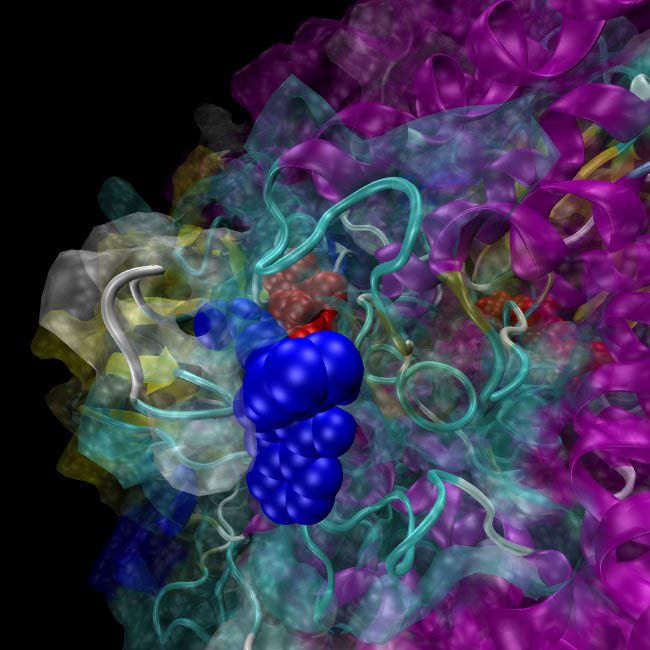
Chemists interested in exploring molecules with therapeutic potential can access libraries that contain thousands of chemical compounds. But even with the help of robotics, physically screening for the promising few to test in the lab can take up to a month. Databases that include millions of commercially available chemicals now allow for faster, vaster and more readily accessible virtual screening.
Biochemists at Southern Methodist University are using this approach — along with supercomputers — to evaluate about 40,000 compounds per day to find the ones that could block a protein that makes chemotherapy drugs less effective. Having combed through 8 million compounds, resulting in a hit list of a few hundred that could plug up the protein, they're now pursuing about 30 of them in the lab.
Predicting Side Effects

When drugs interact with unintended protein targets, they cause side effects, which can include rashes, depression and other unwanted effects. Adverse side effects are the second most common reason (after lack of effectiveness) that potential medicines fail in clinical trials. Predicting the undesired binding events early on could save money and time.
To test whether computer models could identify which drugs were likely to produce adverse side effects, pharmaceutical chemists at the University of California, San Francisco, teamed up with toxicologists at Novartis Institutes for BioMedical Research. They focused on 656 currently prescribed medicines with known safety or side effect records. The scientists used information about thousands of other chemical compounds to predict the drugs' binding to unintended targets — and potential side effects — about half of the time, which represents a big step forward.
Drug Tracking
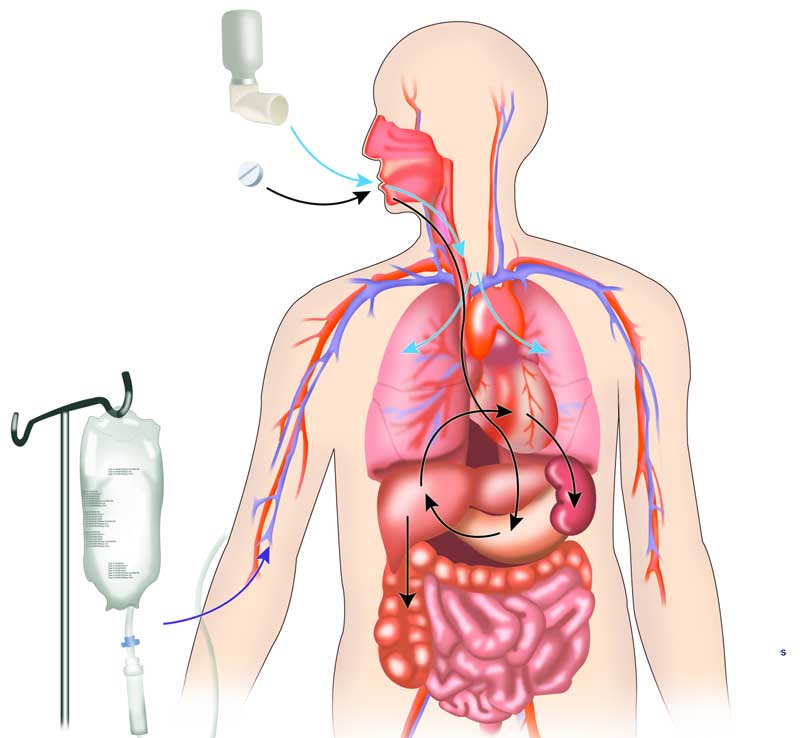
Another way to predict side effects as well as drug efficacy is to figure out how drugs are absorbed, distributed, metabolized and excreted once they enter the body.
To study these pharmacokinetic processes, scientists at the University of Michigan created a computational tool for simulating drug transport at the cellular level. The simulations make it possible to observe and manipulate the distribution of large numbers of drug molecules inside cells and identify which ones are most likely to reach their intended targets. The scientists validate the results by using microscopic imaging to track changes in the distribution of molecules traveling inside the body or cells. The simulations can be used to study and screen drugs already on the market and ones still being tested.
 Live Science Plus
Live Science Plus











