Inside Life Science: What Is an Ontology?
Biomedical researchers face a growing problem in trying to manage their knowledge. As scientists in different disciplines—or even just in different labs—conduct experiments and exchange information, they gather different kinds of data and interpret terms in different ways, sometimes without realizing it.
To make it easier for biologists to understand data and share what they know, the National Institutes of Health is funding computer scientists to build virtual libraries called ontologies. These organize biological knowledge using a universal language.
The problem of too much data
Article continues belowImagine you're a biologist working on, say, brain function in chickens. Before you start your first experiment, you want to find out what research has been done on chicken brains.
First you search the scientific literature—all the journal articles that have been published in your area of interest. Then you tackle the databases.
But even when you pare down the results, you may not be able to interpret or compare them. Your database search may pull up charts from two studies with columns labeled "beak length." The numbers could be averaged, in millimeters, in centimeters, about chicks or roosters, anything. If you don't know what the numbers represent, the data are meaningless to you.
Plus, what you call a chicken may not be what another researcher calls a chicken. This is a rampant problem in gene research, where different scientists call the same DNA segments by different names or use the same names to refer to different segments. If you don't realize that Dr. Smith's data on what he calls a chicken is actually about what you would call an elephant, "you can come up with some really interesting but bogus conclusions," says Karin Remington, who directs the Center for Bioinformatics and Computational Biology at the National Institutes of Health.
Get the world’s most fascinating discoveries delivered straight to your inbox.
Ontologies to the rescue
By establishing a set of official terms, ontologies allow biologists across labs, specialties and countries to share a common vocabulary. Ontology Web Language, a popular choice, gives every protein, every gene, every biological process, a standard name. Everyone will call that beaky, feathered creature that goes bok bok a "chicken," and the term won't be used to describe anything else.
An ontology also establishes what biologists know about the objects they study. For example, a chicken:
- Is a domesticated animal used for food.
- Lays eggs if female.
- Cannot fly long distances.
In the same way, a particular gene may be tagged as "makes proteins that strengthen the cell wall" or "located on chromosome 2."
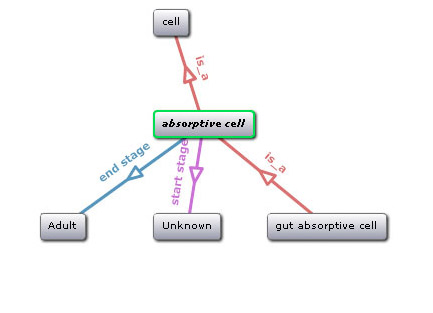
Another benefit is that ontologies organize terms to show how objects and concepts relate to each other. Ontologists may depict these associations as a tree, a flow chart or the nested folder structure on your computer. These visuals make it easier to understand that a chicken is a kind of bird and the cerebellum is part of the brain.
Challenges ahead
As ontologists wrangle all this scientific knowledge into tidy categories, they must clear major hurdles. For instance, researchers don't always agree on terminology. Nor do they necessarily have the same opinion on a protein's function or the connections between certain genes and human diseases. These roles aren't always clear, especially at the cutting edge of discovery. In fact, the sociology of ontology building—how to get communities to develop and agree on standards—is one of the most challenging and rewarding areas of research, says Peter Lyster, also of NIH's Center for Bioinformatics and Computational Biology.
There probably will never be a single, undisputed ontology that contains all scientific knowledge. But that isn't the goal, says Lyster. Instead, it's to develop a series of ontologies that are useful to scientists in specialized fields and that are indexed in one place. It's also to convince scientists around the world that having these ontologies is not only helpful, it's essential.
Learn More:
- National Center for Biomedical Ontology
- Computing Life: How Computation Tools Advance Health and Biology
This Inside Life Science article was provided to LiveScience in cooperation with the National Institute of General Medical Sciences, part of the National Institutes of Health.