Inside Life Science: Studying Protein Shapes Helps Combat HIV
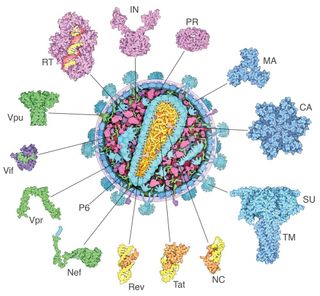
Our bodies contain millions of proteins that carry out a wide range of tasks: delivering oxygen to tissues, defending against infection, digesting food and even making new proteins.
Every protein has a shape that helps the molecule do its job. Collagen in our cartilage and tendons, for instance, has a three-stranded, rope-like structure that makes it strong. Enzymes — proteins that facilitate chemical reactions — typically have a pocket that holds the molecule they act upon. If we can understand protein shape, we can learn more about protein function and how it can contribute to health and disease.
Folding into shape
Proteins are made of long strings of anywhere from 50 to 2,000 amino acids. These basic building blocks link up in a specific sequence dictated by a gene. The chain twists and buckles and, in less than a second, folds into a three-dimensional structure that lets the protein interact with other molecules to trigger all the reactions that run our bodies.
But sometimes things go wrong. If there’s an error in just one amino acid, the protein may misfold and not be able to do its job. Misfolded proteins have been implicated in life-threatening disorders, such as sickle cell disease and cystic fibrosis. If we want to develop drugs that treat or prevent these diseases, we need to know what proteins look like and how to block—or in some cases, boost—their activity.
Solving protein structures can also help pinpoint the Achilles’ heels of disease-causing viruses or bacteria. This structure-based approach has been key in the fight against HIV/AIDS.
Butterfly effect
Sign up for the Live Science daily newsletter now
Get the world’s most fascinating discoveries delivered straight to your inbox.
For the last 25 years, scientists funded by the National Institutes of Health have been studying the structures of proteins involved in HIV. A breakthrough came in 1989, when researchers discovered the shape of HIV protease, an enzyme that helps the virus copy itself and infect more cells. The researchers showed that the protein looks like a butterfly: It is made up of two equal halves with a center that carries out the enzyme’s job. Shutting down this site could stop the virus' spread in the body.
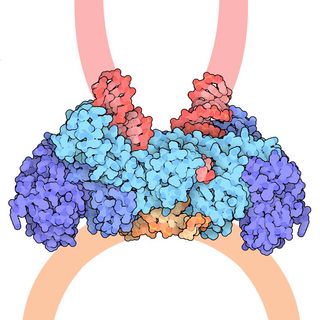
Eventually, other researchers used this knowledge to come up with a new class of drugs, called protease inhibitors, that are now widely used to block the action of HIV protease. As a result, they stop the virus’s spread to new cells.
But HIV is a moving target. When it reproduces inside the body, it can churn out slightly altered versions of itself that may have different protease shapes. This means inhibitor drugs no longer work, allowing the virus to reproduce and spread. Some researchers are now investigating new generations of HIV protease inhibitors that will combat these drug-resistant viral strains.
Citizen scientists
While researchers do the bulk of the work, nonscientists are also getting involved. Through a project called FightAIDS@Home, people can remotely lend their idle home computers to researchers interested in HIV protein structures. The project is part of a growing trend called distributed computing that harnesses the power of personal computers to answer important questions about biology. The typical computers in a scientist's lab can't perform all of the required calculations—but a network of hundreds or even thousands of personal computers can.
FightAIDS@Home, which launched in 2000 and is hosted at the Scripps Research Institute, uses the donated computational power to screen HIV protease structures against potential drug candidates. To date, volunteers have provided about 125,000 years of processing time. This has helped the researchers do more aggressive and ambitious experiments, investigate many more compounds and even offer some general insights into the nature of drug resistance.
All this structural work could lead to new HIV/AIDS treatments that are more potent, are more convenient to take and have fewer side effects.
Learn more:
This Inside Life Science article was provided to LiveScience in cooperation with the National Institute of General Medical Sciences, part of the National Institutes of Health.

